Phytochemicals and Regulation of NF-kB in Inflammatory Bowel Diseases: An Overview of In Vitro and In Vivo Effects.
Study Design
- Tipo de estudio
- Review
- Población
- In vitro and in vivo IBD models
- Intervención
- Phytochemicals and Regulation of NF-kB in Inflammatory Bowel Diseases: An Overview of In Vitro and In Vivo Effects. Various phytochemicals
- Comparador
- None
- Resultado primario
- NF-kB regulation in IBD
- Dirección del efecto
- Positive
- Riesgo de sesgo
- Unclear
Abstract
Inflammatory bowel diseases (IBD) are chronic relapsing idiopathic inflammatory conditions affecting the gastrointestinal tract. They are mainly represented by two forms, ulcerative colitis (UC) and Crohn's disease (CD). IBD can be associated with the activation of nuclear factors, such as nuclear factor-kB (NF-kB), leading to increased transcription of pro-inflammatory mediators that result in diarrhea, abdominal pain, bleeding, and many extra-intestinal manifestations. Phytochemicals can interfere with many inflammation targets, including NF-kB pathways. Thus, this review aimed to investigate the effects of different phytochemicals in the NF-kB pathways in vitro and in vivo models of IBD. Fifty-six phytochemicals were included in this study, such as curcumin, resveratrol, kaempferol, sesamol, pinocembrin, astragalin, oxyberberine, berberine hydrochloride, botulin, taxifolin, naringin, thymol, isobavachalcone, lancemaside A, aesculin, tetrandrine, Ginsenoside Rk3, mangiferin, diosgenin, theanine, tryptanthrin, lycopene, gyngerol, alantolactone, mangostin, ophiopogonin D, fisetin, sinomenine, piperine, oxymatrine, euphol, artesunate, galangin, and nobiletin. The main observed effects related to NF-kB pathways were reductions in tumor necrosis factor-alpha (TNF-α), interleukin (IL)-1β, IL-6, interferon-gamma (IFN-γ), and cyclooxygenase-2 (COX-2), and augmented occludin, claudin-1, zonula occludens-1, and IL-10 expression levels. Moreover, phytochemicals can improve weight loss, stool consistency, and rectal bleeding in IBD. Therefore, phytochemicals can constitute a powerful treatment option for IBD in humans.
TL;DR
The main observed effects related to NF-kB pathways were reductions in tumor necrosis factor-alpha (TNF-α), interleukin (IL)-1β, IL-6, interferon-gamma (IFN-γ), and cyclooxygenase-2 (COX-2) expression levels, and augmented occludin, claudin-1, zonula Occludens- 1, and IL-10 expression levels.
Full Text
metabolites
OH
OH
Review
Phytochemicals and Regulation of NF-kB in Inflammatory Bowel Diseases: An Overview of In Vitro and In Vivo Effects
Lucas Fornari Laurindo 1 , Ana Rita de Oliveira dos Santos 1, Antonelly Cassio Alves de Carvalho 2, Marcelo Dib Bechara 1,2, Elen Landgraf Guiguer 1,2,3 , Ricardo de Alvares Goulart 2, Renata Vargas Sinatora 1,2, Adriano Cressoni Araújo 1,2 and Sandra Maria Barbalho 1,2,3,*
- 1 Department of Biochemistry and Pharmacology, School of Medicine, University of Marília (UNIMAR), Avenida Hygino Muzzy Filho 1001, São Paulo 17525-902, Brazil
- 2 Postgraduate Program in Structural and Functional Interactions in Rehabilitation, University of Marília (UNIMAR), Avenida Hygino Muzzy Filho 1001, São Paulo 17525-902, Brazil
- 3 Department of Biochemistry and Nutrition, School of Food and Technology of Marília (FATEC), Avenida Castro Alves 62, São Paulo 17500-000, Brazil
* Correspondence: [email protected]
Citation: Laurindo, L.F.; Santos, A.R.d.O.d.; Carvalho, A.C.A.d.; Bechara, M.D.; Guiguer, E.L.; Goulart, R.d.A.; Vargas Sinatora, R.; Araújo, A.C.; Barbalho, S.M. Phytochemicals and Regulation of NF-kB in Inflammatory Bowel Diseases: An Overview of In Vitro and In Vivo Effects. Metabolites 2023, 13, 96. https://doi.org/10.3390/metabo13010096
Academic Editor: Marijana Zovko Koncˇic´
Copyright: © 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https:// creativecommons.org/licenses/by/ 4.0/).
Abstract: Inflammatory bowel diseases (IBD) are chronic relapsing idiopathic inflammatory conditions affecting the gastrointestinal tract. They are mainly represented by two forms, ulcerative colitis (UC) and Crohn’s disease (CD). IBD can be associated with the activation of nuclear factors, such as nuclear factor-kB (NF-kB), leading to increased transcription of pro-inflammatory mediators that result in diarrhea, abdominal pain, bleeding, and many extra-intestinal manifestations. Phytochemicals can interfere with many inflammation targets, including NF-kB pathways. Thus, this review aimed to investigate the effects of different phytochemicals in the NF-kB pathways in vitro and in vivo models of IBD. Fifty-six phytochemicals were included in this study, such as curcumin, resveratrol, kaempferol, sesamol, pinocembrin, astragalin, oxyberberine, berberine hydrochloride, botulin, taxifolin, naringin, thymol, isobavachalcone, lancemaside A, aesculin, tetrandrine, Ginsenoside Rk3, mangiferin, diosgenin, theanine, tryptanthrin, lycopene, gyngerol, alantolactone, mangostin, ophiopogonin D, fisetin, sinomenine, piperine, oxymatrine, euphol, artesunate, galangin, and nobiletin. The main observed effects related to NF-kB pathways were reductions in tumor necrosis factoralpha (TNF-α), interleukin (IL)-1β, IL-6, interferon-gamma (IFN-γ), and cyclooxygenase-2 (COX-2), and augmented occludin, claudin-1, zonula occludens-1, and IL-10 expression levels. Moreover, phytochemicals can improve weight loss, stool consistency, and rectal bleeding in IBD. Therefore, phytochemicals can constitute a powerful treatment option for IBD in humans.
Keywords: inflammatory bowel diseases; ulcerative colitis; Crohn’s disease; intestinal inflammation; phytochemicals; nuclear factor kappa B; NF-kB; inflammation; anti-inflammatory
1. Introduction
Inflammatory bowel diseases (IBD) are relapsing idiopathic and chronic inflammatory conditions affecting the gastrointestinal tract. They are mainly characterized by two entities named ulcerative colitis (UC), involving a continuous inflammatory process of the colonic mucosa, and Crohn’s disease (CD), which can produce ulceration lesions anywhere in the gastrointestinal tract. Both environmental and genetic factors are associated with disrupting the intestinal epithelial barrier, resulting in increased permeability and augmented uptake of microorganisms triggering the activation of an imbalanced immune response and leading to inflammation. Environmental factors can be related to diet, food additives, distress, infections, medications, reduced levels of vitamin D, and oxidative stress (OS). Regarding genetic factors, modifications in genes related to the expression of proteins and receptors of immune system cells can be associated with the activation of nuclear factors, such as nuclear Factor-kB (NF-kB). This activation leads to increased transcription of pro-inflammatory
Metabolites 2023, 13, 96. https://doi.org/10.3390/metabo13010096 https://www.mdpi.com/journal/metabolites
mediators, such as interleukin (IL)-1β, IL-6, tumor necrosis factor-α (TNF-α), and interferon (IFN)-γ [1–4].
increased permeability and augmented uptake of microorganisms triggering the activation of an Environmental factors can be related to diet, food additives, distress, infections, medications, reduced levels of vitamin D, and oxidative stress (OS). Regarding genetic factors, modifications in genes related to the expression of proteins and receptors of immune system cells can be associated with the activation of nuclear factors, such as nuclear Factor-kB (NF-kB). This activation leads to increased transcription of proinflammatory mediators, such as interleukin (IL)-1β, IL-6, tumor necrosis factor-α (TNFα), and interferon (IFN)-γ [1–4].
IBD shows a fast-increasing incidence in many countries, and its prevalence is higher in Western regions, specifically Northern Europe and North America. In American adults, the prevalence reaches 1.3%, meaning that 3 million people are affected by this condition. These data make IBD be considered a global burden [5,6]. If it is not adequately treated, its course can severely damage the bowel leading to disability and a profound reduction in the quality of life and capacity to work. Affected patients can suffer from diarrhea, abdominal pain, bleeding, and many extra-intestinal manifestations. Therefore, the therapeutic approach targets the control of inflammation and OS, aiming for mucosal healing and reducing symptoms associated with a reduced risk of hospitalization [7,8]. Corticosteroids, aminosalicylates, antibiotics, immunosuppressive drugs, and monoclonal antibodies are usually used to treat patients. However, these drugs can be associated with several adverse effects, loss of efficacy, and high costs, and many patients are refractory to the treatment. For these reasons, many other substances have been considered to help treat IBD. Several phytochemicals, such as flavonoids, alkaloids, saponins, phenolic acids, and terpenoids, can be effective adjuvants for the therapeutic approach [9–11]. Figure 1 shows the main pathways involved in the pathophysiology of IBD and how the activation of the NF-kB pathways interferes with the occurrence of these diseases.
IBD shows a fast-increasing incidence in many countries, and its prevalence is higher in Western regions, specifically Northern Europe and North America. In American adults, the prevalence reaches 1.3%, meaning that 3 million people are affected by this condition. These data make IBD be considered a global burden [5,6]. If it is not adequately treated, its course can severely damage the bowel leading to disability and a profound reduction in the quality of life and capacity to work. Affected patients can suffer from diarrhea, abdominal pain, bleeding, and many extra-intestinal manifestations. Therefore, the therapeutic approach targets the control of inflammation and OS, aiming for mucosal healing and reducing symptoms associated with a reduced risk of hospitalization [7,8]. Corticosteroids, aminosalicylates, antibiotics, immunosuppressive drugs, and monoclonal antibodies are usually used to treat patients. However, these drugs can be associated with several adverse effects, loss of efficacy, and high costs, and many patients are refractory to the treatment. For these reasons, many other substances have been considered to help treat IBD. Several phytochemicals, such as flavonoids, alkaloids, saponins, phenolic acids, and terpenoids, can be effective adjuvants for the therapeutic approach [9–11]. Figure 1 shows the main pathways involved in the pathophysiology of IBD and how the activation of the NF- κB pathways interferes with the occurrence of these diseases.
Figure 1. Inflammatory bowel diseases (IBD), activation of the nuclear factor-kB (NF-kB), and the possible roles of phytochemicals against this pathway activation during these diseases. ↑, increase;
Phytochemicals can interfere with many targets, including NF-kB pathways [12–15]. Thus, they can be related to regulating many other processes and cascades of inflammatory responses. For these reasons, this review aims to investigate the effects of different phytochemicals in the NF-kB pathways in vitro and in vivo models of IBD. To the best of our knowledge, this is the first review to assess the roles of several phytochemicals in NF-kB regulation in IBD models.
2. Materials and Methods
- 2.1. Focal Question
- 2.2. Language Only studies originally written in English were included in this review.
- 2.3. Databases
- 2.4. Study Selection
- 2.5. Data Extraction
- 2.6. Quality Assessment
The quality assessment of this narrative review was based on the initial literature review of the pathophysiology of UC and CD and activation of NF-kB, as well as its implications for IBD, and, furtherly, on the included in vivo and in vitro studies. The quality assessment was performed by two independent reviewers trained in the Scale for Assessment of Narrative Review Articles (SANRA), a scale of six different items proposed by Baethge et al. [16] that assesses the quality of non-systematic reviews.
Table 1. In vivo and in vitro studies evaluating phytochemicals’ roles as NF-kB regulators on inflammatory bowel diseases.
In Vivo In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-KB-Related Dysregulation Indicators
Table 1. In vivo and in vitro studies evaluating phytochemicals’ roles as NF-kB regulators on inflammatory bowel diseases.
Mechanisms in Regulation of NF-KB in IBD
Related Molecular Mechanisms in Regulation of NF-kB in IBD
DSS-induced FVB/NJ mice model of colitis
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
↓NF-kB-p65-dependent luciferase activity, ↓phosphoNF-kB-p65 and ↓nuclear translocation of p65
Ref.
Phytochemicals
infiltration in the lamina propria, ↑epithelial erosion,
↑TNF-α, ↑IL-6, IL-1β, ↑OS-
3 mg/day of
↑MPO
↑Inflammatory cells infiltration in the lamina propria, ↑epithelial erosion, ↑interstitial edema, and ↓colonic goblet cells
↓NF-kB-p65-dependent luciferase activity, ↓phospho-NF-kB-p65 and ↓nuclear translocation of p65
transgenic mice model of colitis
DSS-induced FVB/NJ mice model of colitis and NFκB-RE-Luc transgenic mice model of colitis
interstitial edema, and ↓colonic goblet cells
↑TNF-α, ↑IL-6, ↑IL-1β, ↑OS-related protein, ↓HO-1, and ↑MPO
3 mg/day of TDNPs for 1 week orally
[17]
Inflammatory cells infiltration and ↑ulcer and granuloma formation
TNBS-induced Wistar Hannover rats model of colitis
NF-kB-related proteins expression and ↓oxidativerelated enzymes expression
↑IL-6, ↑TNF-α, NO
[18]
↓NF-kB-related proteins expression and ↓oxidative-related enzymes expression
↑Inflammatory cells infiltration and ↑ulcer and granuloma formation
TNBS-induced Wistar Hannover rats model of colitis
↑IL-6, ↑TNF-α, ↑MPO, ↑MDA, and ↑NO
20 mg/kg/day for 1 week orally
[18]
↑iNOS, ↑TNF-α, IL-1β, ↑IL-6, ↑nitrite, and ↑S-nitrosylation of IKKβ
100 mg/kg mixed with olive oil in the chow
Body weight, ↑DAI, and ↓colon length
DSS-induced BALB/c mice model of colitis
S-nitrosylation of IKKβ and ↓IκB phosphorylation
[19]
↑iNOS, ↑TNF-α, ↑IL-1β, ↑IL-6, ↑nitrite, and ↑S-nitrosylation of IKKβ
↓S-nitrosylation of IKKβ and ↓IκB phosphorylation
DSS-induced BALB/c mice model of colitis
100 mg/kg mixed with olive oil in the chow
↓Body weight, ↑DAI, and ↓colon length
[19]
Curcumin
Curcumin
↑MPO, NF-kB mRNA, ↑IL-27, ↑TLR4 expression, ↑NF-κB-p65,
TNBS-induced Sprague-Dawley rats model of colitis
↓NF-kB mRNA and ↓NF-kBp65
100 mg/kg/day for 1 week orally
↑DAI score
[20]
TNBS-induced Sprague-Dawley rats model of colitis
↑MPO, ↑NF-kB mRNA, ↑IL-27, ↑TLR4 expression, ↑NF-kB-p65, and ↑IL-27 p28
100 mg/kg/day for 1 week orally ↑DAI score
↓NF-kB mRNA and ↓NF-kB-p65
[20]
↑ diarrhea, thickened
↓Body weight, ↑bloody diarrhea, thickened colon wall, ↓depletion of goblet cells, ↑hemorrhagic intestinal necrosis, ↑mucosal ulcerations, and ↑inflammatory cells infiltration
depletion of goblet cells, ↑hemorrhagic intestinal
↓NF-kB-related proteins expression and ↓oxidative-related enzymes expression
TNBS-induced Sprague-Dawley rats model of colitis
NF-kB-related proteins expression and ↓oxidativerelated enzymes expression
TNBS-induced Sprague-Dawley rats model of colitis
100 mg/kg and 200/mg/kg orally 2 h prior to induction of colitis
200/mg/kg orally 2 h prior to induction of colitis
↑MPO and ↑NF-kB expression
↑MPO and NF-kB expression
[21]
↑mucosal ulcerations, and
inflammatory cells infiltration
↓IFN-γ and IL-12/23p40 genetic expression, ↓phospho-p65-positive expression, ↑IL-10 mRNA, ↓NF-kB-related proteins expression
Bacteria-induced Specific pathogen-free wild-type 129/SvEv mice and germ-free IL 10/mice models of colitis
↑Intestinal-associated ↑crypt hyperplasia, ↑lymphocytic and
Bacteria-induced Specific pathogenfree wild-type 129/SvEv mice and germ-free IL 10/mice
Intestinal-associated crypt hyperplasia,
↓IFN-γ and IL-12/23p40 genetic
0.1, 0.5, and 1% curcumin-supplemented diets for 5 days
↑IL-12/23p40, ↑IFN-γ, ↑NF-kB activation, and ↑pSer276-p65
0.1, 0.5, and 1% curcumin-
[22]
expression, phospho-p65positive expression, ↑IL-10
↑lymphocytic and
↑IL-12/23p40, IFN-γ, ↑NF-Kb activation, and pSer276-p65
neutrophilic infiltrations, and ↑mucosal ulceration
[22]
NF-kB-related proteins expression
diets for 5 days
infiltrations, and
Table 1. Cont.
Related Molecular Mechanisms in Regulation of NF-kB in IBD
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
Ref.
Phytochemicals
↑MPO, ↑LPO and ALP activities; ↑iNOS, and ↑NF-kB-related proteins expression
↓NF-kB-related proteins expression and ↓iNOS expression
25, 50, and 100 mg/kg/day of curcumin orally for 10 days
↑Intestinal ulcers, ↑inflammation in the colon, and ↓colon length
DNCB-induced Wistar rats model of colitis
[23]
↑mRNA do PPARγ, ↑15d-PGJ2, ↑PPAR-γ, ↓COX-2 mRNA, ↓IL-1β mRNA, ↓TNF-α mRNA, ↓IFN-γ mRNA, and ↑IL-4 mRNA
↑IL-1β mRNA, ↑TNF-α mRNA, ↑IFN-γ mRNA, ↑COX-2 mRNA, ↓PPAR-γ,
↑Intestinal epithelial necrosis, ↑glandular destruction, ↑inflammatory cells infiltration, ↓body weight
TNBS-induced Sprague–Dawley rats model of colitis
30 mg/kg/day intraperitoneally for 14 days
[24]
↓PGE2, ↑15d-PGJ2, ↓mRNA IL-4
↓Body weight, intestinal ulcers, ↑inflammatory cells infiltration
↑NF-kB DNA ligation activity, ↑IkB degradation, ↑IL-1β and ↓IL-10
↓NF-kB DNA ligation activity, ↓IkB degradation, and ↓IL-1 β mRNA
TNBS-induced Wistar rats model of colitis
2% of curcumin mixed with the chow for 14 days
[25]
↓Body weight, ↑inflammatory cellular infiltration, and ↑mucosal and muscle damage
↑MPO, ↑IL-1 β, ↑NF-kB DNA ligation activity, and ↑p38 MAPK
↓NF-kB DNA ligation activity and ↓p38 MAPK
TNBS-induced BALB/c mice model of colitis
0.25% of curcumin mixed with the chow for 10 days
[26]
↑Hemorrhagic and ulcerative damage to the distal colon, ↑mucosal congestion, ↑leucocyte cellular infiltrate in the submucosa, and ↓body weight
↓Serine protease activities, ↓IFN-γ and IL-12 p40 mRNA levels, ↓NF-kB-related proteins expression
↑NO, ↑MPO, ↑MDA, ↑protease activities, ↑IFN-γ and IL-12 p40 mRNAs, and ↑iNOS
TNBS-induced C3H mice model of colitis
25–300 mg/kg/day of curcumin orally for 10 days
[27]
↑p65 nuclear expression, ↑IkB degradation, ↑macrophage infiltration, ↑IL-18, ↑NF-kB DNA ligation activity, ↑IL-6 mRNA, ↑IFN-γ mRNA, ↑TNF-α mRNA, and ↑IL-12 mRNA
↓IkB degradation, ↓NF-kB DNA ligation activity, ↓IL-6 mRNA, ↓IFN-γ mRNA, ↓TNF-α mRNA, and ↓IL-12 mRNA
↓Body weight, ↑crypts distortion, ↓goblet cells, and mononuclear cells infiltration
TNBS-induced C57BL/6 and BALB/c mice models of colitis
0.5%, 2.0%, or 5.0% of curcumin mixed with the chow for 7 days
[28]
BALB/c mice model mixed with the infiltration, and ligation activity, and ↑p38
[26]
and ↓p38 MAPK
Table 1. Cont.
ulcerative damage to the distal colon, ↑mucosal congestion,
Related Molecular Mechanisms in Regulation of NF-kB in IBD
↓Serine protease activities, ↓IFN-γ and IL-12 p40 mRNA
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
↑NO, ↑MPO, protease activities, ↑IFN-γ and IL-12 p40 mRNAs, and ↑iNOS
, 96 6 of 44
25–300 mg/kg/day of curcumin orally for 10 days
Ref.
Phytochemicals
TNBS-induced C3H
[27]
NF-kB-related proteins expression
infiltrate in the submucosa, and ↓body
Weight loss, diarrhea, bloody stool, ↑MPO activity, ↑DAI score, ↑colonic cytokine levels, and ↑visceral pain
10 µL 4.5 mM and 10 µL 45 mM/day intraperitoneally for 4 days
↑pNF-kB, ↑TNF-α mRNA, ↑TNF-α, ↑TGF-β mRNA, ↑TGF-
and PGE expression mRNA, and IkBα phosphorylation and
↓pNF-kB, ↓TNF-α mRNA, and ↓TGF-β mRNA
TNBS-induced C57BL/6 mice model of colitis
[29]
↑p65 nuclear expression, degradation, ↑macrophage
↓Body weight, ↑crypts
↓IkB degradation, ↓NF-kB DNA ligation activity, ↓IL-6 mRNA, ↓IFN-γ mRNA, ↓TNFα mRNA, and IL-12 mRNA
TNBS-induced
0.5%, 2.0%, or 5.0%
Colon inflammation (lymphocyte infiltration and distortion of glands), weight loss,
↓p65 nuclear translocation, ↓IKK phosphorylation, ↓COX-2 mRNA, and ↓IkBα phosphorylation and degradation
goblet cells, and mononuclear cells
↑Colon inflammation measured by COX-2 and PGE2 expression levels
[28]
DNA ligation activity, ↑IL-6 mRNA, ↑IFN-γ mRNA, ↑TNF-
↑p65 nuclear translocation, ↑COX-2, ↑PGE2
10–50 µM during 1 h of incubation
↑IL-6, ↑IL-1β, IFN-γ, ↑TNF-α, ↑p-IkBα, ↓SIRT1
mice models of colitis
with the chow for 7 days
and 100 mg/kg on alternate days
[30]
LPS-treated Caco-2 cells
↓p-IkBα and SIRT1 [31]
C57BL/6 mice model
Weight loss, diarrhea, bloody stool, ↑MPO activity, DAI score, ↑colonic cytokine levels, and ↑visceral pain
inflammatory cytokines
Colon inflammation (lymphocyte infiltration and distortion of glands), weight loss, and ↑serum pro-inflammatory cytokines
10 µL 4.5 mM and 10 µL 45 mM/day intraperitoneally for 4 days
Resveratrol TNBS-induced
↑NF-kB-DNA binding complex, ↑IKKβ catalytic activity, ↑ERK phosphorylation, expression, ↑STAT3
Resveratrol
100 µL of 10, 50, and 100 mg/kg on alternate days orally for 7 days
↑pNF-κB, ↑TNF-α mRNA, ↑TNF-α, ↑TGF-β mRNA, ↑TGF-
↓pNF-Κb, TNF-α mRNA, and ↓TGF-β mRNA
DSS-induced C57BL/6 mice model of colitis
↑IL-6, ↑IL-1β, ↑IFN-γ, ↑TNF-α, ↑p-IkBα, ↓SIRT1 ↓p-IkBα and ↑SIRT1 [31]
↓ERK phosphorylation, ↓NFkB-DNA binding complex, and
↑Histopathological score
[29]
C57BL/6 mice model
DSS-induced ICR 10 mg/kg/day orally for 7 days
↑NF-kB-DNA binding complex, ↑IKKβ catalytic activity, ↑ERK phosphorylation, ↑iNOS expression, ↑STAT3
↓p65 nuclear translocation, ↓IKK phosphorylation, ↓COX-2
p65 nuclear translocation, COX-2, PGE
↓ERK phosphorylation, ↓NF-kB-DNA binding complex, and ↓IKKβ catalytic activity
[30]
DSS-induced ICR mice model of colitis
10 mg/kg/day orally for 7 days ↑Histopathological score
cells h of incubation measured by COX-2
inflammation, presence of adhesions between
[32]
bowel and other organs, ulcers, crypt
↑Macroscopic inflammation, presence of adhesions between the colon and small bowel and other organs, ulcers, crypt distortion, ↑leukocyte involvement, ↑pro-inflammatory cytokines production, and weight loss
↑MPO, ↑TNF-α, ↑COX-2, ↑PGD2
leukocyte involvement, ↑proinflammatory cytokines production, and weight
TNBS-induced Wistar mice model of colitis
10 mg/kg/day orally for 14 days
↑MPO, ↑TNF-α, ↑NF-kB p65, ↑COX-2, ↑PGD2
↑Inflammatory mucosa cells apoptosis and ↓NF-kB p65
[33]
DSS-induced colitis in antibiotics-treated pseudo-germ-free mice and LPS-stimulated RAW264.7 cells
3-(4-Hydroxyphenyl)-propionic acid in antibiotics-treated pseudo-germ-free mice and LPSstimulated RAW264.7 cells
3-(4-Hydroxyphenyl)-propionic acid
↑Intestinal inflammation and ↑OS both in vivo and in vitro
↑NF-kB-related activation proteins and MAPK
↓NF-kB-related activation proteins and MAPK
↑NF-kB-related activation proteins and ↑MAPK
↓NF-kB-related activation proteins and ↓MAPK
inflammation and ↑OS and in vitro
[34]
-
[34]
orally for 14 days injury, and altered gut MCP-1 mRNA, iNOS mRNA,mRNA, IL-10 mRNA, TLR4,
production, and weight
Table 1. Cont.
3-(4-Hydroxyphenyl)-propionic acid in antibiotics-treated pseudo-germ-free mice and LPSstimulated
↑Intestinal
DSS-induced colitis in antibiotics-treated pseudo-germ-free mice and LPSstimulated
↑NF-kB-related activation proteins and
↓NF-kB-related activation proteins and
Related Molecular Mechanisms in Regulation of NF-kB in IBD
[34]
↑Intestinal inflammation and ↑OS
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
both in vivo in vitro
↑NF-kB-related activation proteins and
↓NF-kB-related activation proteins and MAPK
Ref.
Phytochemicals
[34]
COX-2 mRNA, ↓iNOS mRNA, ↓IL-6 mRNA, IL-1β mRNA, ↓TNF-α mRNA, ↓TLR4 mRNA, p-NF-κB/NF-κB ratio
↓COX-2 mRNA, ↓iNOS mRNA, ↓IL-6 mRNA, ↓IL-1β mRNA, ↓TNF-α mRNA, ↓TLR4 mRNA, and ↑p-NF-kB/NF-kB ratio
Sesamol
Sesamol
DAI, histopathological changes, and ↓intestinal barrier integrity
↑DAI, histopathological changes, and ↓intestinal barrier integrity
↑COX-2, ↑iNOS, IL-6, ↑IL-1β, ↑TNF-α, ↑TLR4
100 mg/kg/day orally for 6 weeks
↑COX-2, ↑iNOS, ↑IL-6, ↑IL-1β, ↑TNF-α, ↑TLR4
DSS-induced C57BL/6 mice model of colitis
100 mg/kg/day orally for 6 weeks
C57BL/6 mice model of colitis
[35]
↓COX-2 mRNA, ↓iNOS mRNA, ↓IL-6 mRNA, IL-1β mRNA,
[35]
Sesamol DSS-induced
DAI, histopathological
↑COX-2, IL-6, ↑IL-1β,
100 mg/kg/day
↓intestinal barrier integrity
[35]
↓TLR4 mRNA, and ↑p-NF-κB/NF-κB ratio
↑IL-6, ↑IL-1β, ↑TNF-α, ↑IL-1β IL-6 mRNA, ↑TNF-a COX-2 mRNA, MCP-1 mRNA, iNOS mRNA, ↓IL-10 mRNA, TLR4, ↑NLRP3, ↑MAPK1, ↑NF-kB-related
IL-1β mRNA, ↓IL-6 mRNA, ↓TNF-a mRNA, mRNA, ↓MCP-1 mRNA, ↓iNOS mRNA, ↑IL-10 mRNA, ↓TLR4, ↓NLRP3, MAPK1, ↓MyD88, ↓p-NF-kB-P65
of colitis
↑IL-6, ↑IL-1β, ↑TNF-α, ↑IL-1β mRNA, ↑IL-6 mRNA, ↑TNF-a mRNA, ↑COX-2 mRNA, ↑MCP-1 mRNA, ↑iNOS mRNA, ↓IL-10 mRNA, ↑TLR4, ↑NLRP3, ↑MAPK1, ↑NF-kB-related proteins expression, ↓ZO-1, ↓occludin, and ↓claudin-1
TNF-α, ↑IL-1β
IL-1β mRNA, IL-6 mRNA, ↓TNF-a mRNA, ↓COX-2 mRNA, MCP-1 mRNA, ↓iNOS mRNA, ↑IL-10 mRNA, ↓TLR4, ↓NLRP3, MAPK1, ↓MyD88, ↓p-NF-kB-P65
IL-1β mRNA, ↓IL-6 mRNA,
Kaempferol
Kaempferol
[36]
IL-6 mRNA, ↑TNF-a mRNA, ↑COX-2 mRNA,
↓TNF-a mRNA, ↓COX-2 mRNA, ↓MCP-1 mRNA, ↓iNOS mRNA, ↑IL-10
injury, and altered gut microbiota
↑DAI, ↓colon length, ↑intestinal mucosal injury, and altered gut microbiota
of colitis
↑intestinal mucosal injury, and altered gut
50 mg/kg/day orally for 14 days
DSS-induced C57BL/6 mice [36] model of colitis
50 mg/kg/day orally for 14 days
13, 96 7 of 44
C57BL/6 mice model of colitis
[36]
↑MCP-1 mRNA, iNOS mRNA, ↓IL-10 mRNA, TLR4, ↑NLRP3,
mRNA, ↓TLR4, ↓NLRP3, ↓MAPK1, ↓MyD88, ↓p-NF-kB-P65
NF-kB-related
proteins expression, ↓ZO-1, ↓occludin, and ↓claudin-1
↓TLR4 mRNA, ↑ZO-1 mRNA, ↑occludin mRNA,
TLR4 mRNA, ↑ZO-1 mRNA, ↑occludin mRNA, ↑Muc2
↑TLR4 mRNA,↑MCP-1 mRNA, ↑IL-1β mRNA, ↑TNF-α mRNA, ↑COX-2
MCP-1 mRNA, ↑IL-1β mRNA, TNF-α mRNA, ↑COX-2 mRNA, IFN-γ mRNA, ↑p-IκBα, p-IKKα/β,
↑DAI, ↑intestinal mucosal injury, ↑inflammatory cells infiltration, and ↓colon length
↑Muc2 mRNA, ↓p-IκBα, ↓p-IKKα/β, and ↓p-p65, ↓MCP-1 mRNA, ↓IL-1β
mucosal injury, ↑inflammatory cells infiltration, and ↓colon
200 µL of 50, 75, and 100 mg/kg/day orally for 7 days
mRNA, ↓p-IκBα, ↓p-IKKα/β, and ↓p-p65, MCP-1 mRNA, ↓IL-1β mRNA, ↓TNF-α mRNA,
and 100 mg/kg/day orally for 7 days
DSS-induced C57BL/6 mice model of colitis
[37]
C57BL/6 mice model of colitis
[37]
mRNA, ↑IFN-γ mRNA, ↑p-IκBα, ↑p-IKKα/β, and ↑p-p65
mRNA, ↓TNF-α mRNA, ↓COX-2 mRNA, ↓IFN-γ mRNA
Astragalin
Astragalin
COX-2 mRNA, ↓IFN-γ mRNA
↑Pro-inflammatory cytokines production colon cells proliferation in vitro and ↓colon length, ↑pro-inflammatory cytokines production, and weight loss in vivo
↑Cells proliferation, ↑TNF-α mRNA, IL-8 mRNA, ↑IL-6 mRNA, ↑IκBα phosphorylation, and ↑NF-kBDNA binding in vitro and ↑IκBα phosphorylation, ↑TNFα mRNA, ↑IL-8 mRNA, and ↑IL-6 mRNA in vivo
↓Cells proliferation, ↓TNF-α mRNA, IL-8 mRNA, ↓IL-6 mRNA, phosphorylation, and ↓NF-kBDNA binding in vitro and ↓IκBα phosphorylation, ↓TNFα mRNA, ↓IL-8 mRNA, and ↓IL-6 mRNA in vivo
TNF-α -stimulated
↑Cells proliferation, ↑TNF-α mRNA, ↑IL-8 mRNA, ↑IL-6 mRNA, ↑IκBα phosphorylation, and ↑NF-kB-DNA binding in vitro and ↑IκBα phosphorylation, ↑TNF-α mRNA, ↑IL-8 mRNA, and ↑IL-6 mRNA in vivo
↓Cells proliferation, ↓TNF-α mRNA, ↓IL-8 mRNA, ↓IL-6 mRNA, ↓IκBα phosphorylation, and ↓NF-kB-DNA binding in vitro and ↓IκBα phosphorylation, ↓TNF-α mRNA, ↓IL-8 mRNA, and ↓IL-6 mRNA in vivo
↑Pro-inflammatory cytokines production and ↑colon cells proliferation in vitro and ↓colon length, ↑pro-inflammatory cytokines production, and weight loss in vivo
human colonic epithelial cells in vitro and DSSinduced C57BL/6 mice model of colitis
and 100 µM incubated for 24 h in vitro and 2 and 5 mg/kg/day orally
TNF-α -stimulated HCT-116 and HT-29 human colonic epithelial cells in vitro and DSS-induced C57BL/6 mice model of colitis in vivo
0, 20, 40, 60, 80, and 100 µM incubated for 24 h in vitro and 2 and 5 mg/kg/day orally for 7 days in vivo
[38]
[38]
↑Inflammation in vitro
Metabolites 2023, 13, 96 8 of 47
↑Pro-inflammatory
↑Cells proliferation, ↑TNF-α
↓Cells proliferation, ↓TNF-α
TNF-α -stimulated
Astragalin length and ↑p-p65
↓ ↓
Table 1. Cont.
HCT-116 and HT-29 human colonic
epithelial cells vitro and DSSinduced C57BL/6 mice model of colitis
[38]
and ↓colon length,
and ↑colon cells
mRNA, ↑IκBα
mRNA, ↓IκBα phosphorylation, and ↓NF-kBDNA binding in vitro and ↓IκBα phosphorylation, ↓TNFα mRNA, ↓IL-8 mRNA, and ↓IL-6 mRNA in vivo
vitro
in vitro
DNA binding in vitro and
DNA binding in vitro and
and 100 µM
and ↓colon length, ↑pro-inflammatory cytokines production,
in vitro and 2 and 5 mg/kg/day orally for 7 days in vivo
DNA binding in vitro and ↑IκBα phosphorylation, ↑TNFα mRNA, ↑IL-8 mRNA, and
DNA binding in vitro and ↓IκBα phosphorylation, ↓TNFα mRNA, ↓IL-8 mRNA, and
Related Molecular Mechanisms in Regulation of NF-kB in IBD
IκBα phosphorylation, ↓TNFα mRNA, IL-8 mRNA, and ↓IL-6 mRNA in vivo
[38]
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
α mRNA, ↑IL-8 mRNA, and ↑IL-6 mRNA in vivo
and ↓colon length, ↑pro-inflammatory cytokines production, and weight loss in vivo
mice model of colitis in vivo
for 7 days in vivo
Ref.
Phytochemicals
vitro and DSSinduced C57BL/6 mice model of colitis
and 2 and 5 mg/kg/day orally for 7 days in vivo
DNA binding in vitro and ↑IκBα phosphorylation, ↑TNFα mRNA, ↑IL-8 mRNA, and ↑IL-6 mRNA in vivo
in vivo
↑ in vitro
↑ and weight loss,
↓
↑iNOS, ↑IFNγ, ↑IL-6, ↑ TLR4, ↑p65
↓Pro-inflammatory cytokines expression, ↓NF-kB-luciferase activity
↑TNF-α, ↑COX-2, ↑iNOS, ↑IFN-γ, ↑IL-6, ↑IL-15, ↑TLR4, ↑p65 phosphorylation, ↑IκBα phosphorylation, and ↓NO in vitro and ↑p65 phosphorylation, ↑TLR4 mRNA, ↑Myd88 mRNA,
↓Pro-inflammatory cytokines expression, NF-κB-luciferase activity and TLR4/MD2 · LPS interaction in vitro and ↓TLR4 mRNA, Myd88 mRNA, ↓iNOS mRNA, ↓COX-2 mRNA, ↓TNFα mRNA, and p65 phosphorylation in vivo
↑TNF-α, COX-2, ↑iNOS, ↑IFNγ, ↑IL-6, ↑IL-15, TLR4, ↑p65 phosphorylation, ↑IκBα phosphorylation, and ↓NO in
↑Inflammation in vitro and weight loss, ↑intestinal tissue damage, ↑mucosa muscle thickness, ↑neutrophil infiltration, ↑diarrhea, ↑microbiota alterations, and ↑blood in stool in vivo
NF-κ
Inflammation in vitro and weight loss, ↑intestinal tissue
Pinocembrin
Pinocembrin
intestinal tissue damage, ↑mucosa muscle thickness, ↑neutrophil infiltration,
↑IκBα phosphorylation, and ↓ in vitro and ↑
↑TNF-α, ↑COX-2, ↑iNOS, ↑IFNγ, ↑IL-6, ↑IL-15, TLR4, ↑p65 phosphorylation, ↑IκBα phosphorylation, and ↓NO in
incubated for 24 h in vitro and 100 mg/kg/day orally
Pro-inflammatory cytokines expression, ↓NF-κB-luciferase activity and TLR4/MD2 · LPS interaction in vitro and ↓TLR4 mRNA, ↓Myd88 mRNA, ↓iNOS mRNA, ↓COX-2 mRNA, ↓TNFα mRNA, and ↓p65 phosphorylation in vivo
RAW264.7 and Caco2 cells in vitro and DSS-induced C57BL/6 mice model
incubated for 24 h in vitro and 25, 50, and 100 mg/kg/day orally
↑
and ↓TLR4/MD2 · LPS interaction in vitro and
LPS-stimulated RAW264.7 and Caco-2 cells in vitro and DSS-induced C57BL/6 mice model of colitis in vivo
0–200 µM incubated for 24 h in vitro and 25, 50, and 100 mg/kg/day orally for 9 days in vivo
in vitro and ↓ mRNA, ↓Myd88 mRNA, ↓ mRNA, ↓COX-2 mRNA, ↓TNFα p65 phosphorylation in vivo
in vitro and DSS-induced C57BL/6 mice model
LPS-stimulated RAW264.7 and Caco2 cells in vitro and
0–200 µM incubated for 24 h in vitro and 25, 50,
Pinocembrin
[39]
[39]
↓TLR4 mRNA, ↓Myd88 mRNA, ↓iNOS mRNA, ↓COX-2 mRNA, ↓TNF-α
vitro and ↑p65 phosphorylation, ↑TLR4 mRNA, Myd88 mRNA, ↑iNOS mRNA, COX-2 mRNA, ↑
↑
damage, mucosa muscle thickness,
↑TLR4 mRNA, Myd88 mRNA, ↑ COX-2 mRNA, ↑
[39]
microbiota alterations, and ↑blood in stool in vivo
vitro and ↑p65 phosphorylation, ↑ Myd88 mRNA, ↑iNOS mRNA, ↑COX-2 mRNA,
↑
neutrophil infiltration,
mRNA, and ↓p65 phosphorylation in vivo
↑iNOS mRNA, ↑COX-2 mRNA, ↑TNF-α mRNA
of colitis in vivo
for 9 days in vivo
↑
↓ ↑ ↑ZO-2, ↑occludin, claudin-1
↓MPO, ↑ZO-1, ↑ZO-2, ↑occludin, ↑JAM-A, and ↑claudin-1 expressions, ↓IL-6, ↓IL-1β, ↓IL-17, ↓TNF-α, and IFN-γ expressions, ↑p65 (cytoplasm), ↓p65 (nucleus), ↓p-IκBα/IκBα ratio, ↓TLR4, ↓MyD88, ↓NF-kB-p65 translocation, ↓IκBα phosphorylation
ZO-2, ↑occludin,
Oxyberberine
JAM-A, and claudin-1 expressions, IL-6, ↓IL-1β, ↓IL-17, ↓TNF-α, and IFN-γ expressions, ↑p65 (cytoplasm), ↓p65 (nucleus), ↓p-IκBα/IκBα ratio, ↓TLR4, ↓MyD88, ↓NF-κB-p65
Oxyberberine
ZO-2,
↑MPO, ↓ZO-1, ↓ZO-2, ↓occludin, ↓JAM-A, ↓claudin-1, ↑IL-6, ↑IL-1β, ↑IL-17, ↑TNF-α, ↑IFN-γ, ↑TLR4, ↑MyD88, ↑p-IκBα, ↑p65 (nucleus), ↓IκBα, ↓p65 (cytoplasm)
ZO-2, ↓occludin, ↓JAM-A, ↓claudin-1, ↑IL-6, ↑IL-1β, ↑IL-17, ↑TNF-α, ↑IFN-γ, ↑TLR4, MyD88, ↑p-
IL-6, ↓IL-1β, ↓IL-17, ↓TNF-α γ ↑ ↓p65 (nucleus), ↓p-IκBα κBα ↓TLR4, ↓MyD88, ↓NF-κ
↓MPO, ↑ ZO-2, ↑occludin, ↑JAM-A, and claudin-1 expressions, IL-6, ↓IL-1β, ↓IL-17, ↓TNF-α, and IFN-γ expressions, ↑p65 (cytoplasm), ↓p65 (nucleus),
occludin, ↓ ↑IL-6, ↑IL-1β, ↑IL-17, ↑TNF-α, ↑IFN-γ, ↑TLR4, MyD88, ↑
Shaggy hair, low vitality, body weight loss, diarrhea, occult fecal blood, and ↑DAI
Shaggy hair, low vitality, body weight loss, diarrhea, occult fecal blood, and ↑DAI
Oxyberberine
12.5, 25, and 50 mg/kg/day orally/7 days
↑MPO, ↓ZO-1, ZO-2, ↓occludin, ↓JAM-A, ↓claudin-1,
DSS-induced BALB/c mice model of colitis
12.5, 25, and 50 mg/kg/day orally/7 days
DSS-induced BALB/c mice model of colitis
mg/kg/day
[40]
mice model of colitis
Shaggy hair, low vitality, body weight
12.5, 25, and 50
↑
p65 (nucleus), ↓IκBα,
IL-17, ↑TNF-α,
p65 (nucleus), ↓IκBα,
[40]
MyD88, ↑pp65 (nucleus), IκBα,
p-IκBα/IκBα ratio, ↓TLR4,
Bacterial metabolite
IκBα
IκBα
Bacterial metabolite
NF-κB-p65 translocation, ↓IκBα
↓
↓IL-1 mRNA, ↓IL-1β mRNA,
Bacterial metabolite
Weight loss, ↓survival
↓IL-1 mRNA, ↓IL-1β mRNA,
↑ ↑ IL-6 mRNA, ↑TNF-α, ↑IFN-γ mRNA, ↓IL-4 mRNA, ↓IL-10 mRNA, ↑iNOS, ↑MPO, p-NF-κ
↓IL-1 mRNA, ↓IL-1β mRNA, ↓IL-6 mRNA, ↓IL-12 mRNA, ↓TNF-α, ↓IFN-γ mRNA, ↑IL-4 mRNA, ↑IL-10 mRNA, ↓activity of iNOS, MPO,
IL-12 mRNA, ↓TNF-α, IFN-γ mRNA, ↑IL-4 mRNA, IL-10 mRNA, ↓ MDA, ↑
colon length, colon weight, ↑DAI, daily activity,
IL-12 mRNA, ↓TNF-α, IFN-γ mRNA, ↑IL-4 mRNA, IL-10 mRNA, ↓activity of iNOS, MPO, and MDA, ↓p-NF-κB, ↑p-STAT3
↑IL-6 mRNA, ↑IL-12 mRNA, ↑TNF-α, ↑IFN-γ mRNA, ↓IL-4 mRNA, ↓IL-10 mRNA, ↑iNOS, ↑MPO, ↑ p-NF-κB
Weight loss, ↓survival rate, ↓colon length, ↓colon weight, ↑DAI, ↓daily activity, anorexia, ↑inflammatory cells infiltration, ↑intestinal edema, and ↑microscopic damage scores
↑DAI,
↓IL-1 mRNA, IL-1β mRNA, ↓IL-6 mRNA, IL-12 mRNA, ↓TNF-α, ↓IFN-γ mRNA, ↑IL-4
survival
↑IL-1 mRNA, ↑IL-1β mRNA, ↑IL-6 mRNA, ↑IL-12 mRNA, ↑TNF-α, ↑IFN-γ mRNA, ↓IL-4
Berberine hydrochloride
↑IL-1 mRNA, ↑IL-1β mRNA, ↑IL-6 mRNA, IL-12 mRNA, ↑TNF-α, IFN-γ mRNA, ↓IL-4
Berberine hydrochloride
[41]
colon length, ↓colon weight, ↑DAI, ↓daily activity,
orally/6 weeks
DSS-induced Wistar mice model of colitis
DSS-induced Wistar mice [41] model of colitis
10, 30, and 50 mg/kg/day orally/6 weeks
anorexia, inflammatory cells
mg/kg/day orally/6 weeks
[41]
↑inflammatory cells
mRNA, IL-10 mRNA, ↓activity of iNOS, MPO, and MDA, ↓p-NF-κB, ↑p-STAT3
mRNA, IL-10 mRNA, ↑iNOS, ↑MPO, ↑MDA, p-NF-κB
mRNA, ↓IL-10 mRNA, ↑iNOS, ↑MPO, ↑MDA, ↑p-NF-kB
and MDA, ↓p-NF-kB, ↑p-STAT3 expression, ↑ZO-1 mRNA, ↑VCAM-1
inflammatory cells
mRNA, ↑occludin mRNA, and ↑claudin-1 mRNA
Metabolites 2023, 13, 96
,
Metabolites 2023, 13, 96 8 of 44
Table 1. Cont.
↑
expression, ↑
infiltration, ↑intestinal
expression, ↑ZO-1 mRNA, occludin mRNA, and claudin-1 mRNA
↑VCAM-1 mRNA, ↑occludin mRNA, and claudin-1 mRNA
edema, and ↑microscopic damage scores
↑
Related Molecular Mechanisms in Regulation of NF-kB in IBD
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
↑
↑microscopic damage scores
Ref.
Phytochemicals
↓Lipid peroxidation,
↓Lipid peroxidation,
↑antioxidant SOD, and CAT
↓Lipid peroxidation, ↑antioxidant SOD, and CAT expressions, ↓pro-inflammatory cytokines TNF-α, IL-1β, and IL-6 expressions, ↑IL-10 expression, ↓iNOS, and ↓COX-2 activities, ↓TLR4 expression, and ↓NF-kB activation (phosphorylation and nuclear translocation)
antioxidant SOD, and CAT expressions, pro-inflammatory cytokines TNF-α, IL-1β, and IL6 expressions, IL-10 expression, iNOS, and ↓COX-2 activities, TLR4 expression, and ↓NF-kB activation (phosphorylation and nuclear
pro-inflammatory cytokines TNF-α, IL-1β, and IL6 expressions, IL-10 expression, iNOS, and ↓COX-2 activities, TLR4 expression, and ↓NF-kB activation (phosphorylation and nuclear
Lipid peroxidation, ↓SOD, ↓CAT, TNF-α, IL-1β, ↑IL-6, ↓IL-10, ↑ ↑TLR4, and ↑NF-kB activation (phosphorylation and nuclear
Lipid peroxidation, ↓SOD, ↓CAT, TNF-α, IL-1β, ↑IL-6, ↓IL-10, ↑iNOS, COX-2, ↑TLR4, and ↑NF-kB activation (phosphorylation and nuclear
expressions, pro-inflammatory cytokines TNF-α, IL-1β, and IL6 expressions, expression, ↓
↑Lipid peroxidation, ↓SOD, ↓CAT, ↑TNF-α, ↑IL-1β, ↑IL-6, ↓IL-10, ↑iNOS, ↑COX-2, ↑TLR4, and ↑NF-kB activation
Berberine
IL-1β, ↑IL-6, ↓IL-10, ↑iNOS, COX-2, ↑TLR4, and ↑NF-kB activation (phosphorylation and nuclear translocation)
Berberine
Intestinal inflammation measured by shortened, thickened, and erythematous colon
C3H/HeN and C3H/HeJ mice models of colitis
measured by shortened, thickened, and erythematous colon
Intestinal inflammation measured by shortened, thickened, and erythematous colon
dissolved in 2% Tween 80 solution/day
TNBS-induced C3H/HeN and C3H/HeJ mice models of colitis
10 and 20 mg dissolved in 2% Tween 80 solution/day orally/5 days
Tween 80 solution/day orally/5 days
[42]
C3H/HeN and C3H/HeJ mice models of colitis
[42]
[42]
(phosphorylation and nuclear translocation)
and ↓NF-kB activation (phosphorylation and nuclear
↓MPO activity, ↓pro-
↓ ↓
↓pro-
↓MPO activity, ↓pro-inflammatory cytokines IL-6, IL-1β, IL-12, IL-2, and TNF-α expressions, ↑IL-10 expression, ↑antioxidant enzymes SOD, CAT, and GSH-Px expressions, ↓MDA expression, ↓p65 phosphorylation, ↓IκBα phosphorylation, and ↑IκBα
inflammatory cytokines IL-6, IL-1β, IL-12, IL-2, and TNF-α expressions, IL-10 expression,
inflammatory cytokines IL-6, IL-1β, IL-12, IL-2, and TNF-α expressions, IL-10 expression,
Eriodictyol
Eriodictyol
MPO, IL-6, IL-1β, ↑IL 12, ↑IL-2, ↑TNF-α, IL-10, ↓SOD, ↓CAT, ↓GSH-Px, MDA, ↑TLR4, p-p65, and
α
Weight loss, colon crypt destruction, mucosal ulceration, and colon inflammatory cells
↑IL-2, ↑TNF-α, IL-10, ↓SOD, ↓CAT, ↓GSH-Px, MDA, ↑TLR4, ↑p-IκBα, p-p65, and ↓IκBα
destruction, mucosal ulceration, and colon inflammatory cells
↑MPO, ↑IL-6, ↑IL-1β, ↑IL 12, ↑IL-2, ↑TNF-α, ↓IL-10, ↓SOD, ↓CAT, ↓GSH-Px, ↑MDA, ↑TLR4, ↑p-IκBα, ↑p-p65, and ↓IκBα
Weight loss, colon crypt destruction, mucosal ulceration, and colon inflammatory cells infiltration
↑ ↑TNF-α, IL-10, ↓SOD, ↓CAT, ↓ ↑TLR4,
expressions, ↑
↑antioxidant enzymes SOD, CAT, and GSH-Px expressions,
Wistar mice models of colitis
mg/kg/day orally/7 days
TNBS-induced Wistar mice models of colitis
5, 20, and 50 mg/kg/day orally/7 days
↑antioxidant enzymes SOD, CAT, and GSH-Px expressions,
Wistar mice models of colitis
mg/kg/day orally/7 days
[43]
[43]
↓MDA expression, ↓p65 phosphorylation, ↓IκBα phosphorylation, and ↑IκBα
↓ ↓ phosphorylation, ↓IκBα
↓MDA expression, ↓p65 phosphorylation, ↓IκBα phosphorylation, and ↑IκBα
↑IκBα
Diffuse necrosis, congestion, and hemorrhage of the mucosal layer and submucosal edema, congestion, and immune/inflammatory
↓ ↓TLR4 content, ↓CD68 cells infiltration, ↓IL-6 content, NF-κB expression,
Betulin
congestion, and hemorrhage of the mucosal layer and submucosal edema, congestion, and immune/inflammatory cells infiltration
Diffuse necrosis, congestion, and hemorrhage of the mucosal layer and submucosal edema, congestion, and immune/inflammatory cells infiltration
↓LDH activity, ↓TLR4 content, ↓CD68 cells infiltration, ↓IL-6 content, ↓NF-kB expression, ↓TNF-α expression, ↓IL-1β, ↓caspase-3 expression, and ↓caspase-8 expression
Betulin
↑CRP, ↑LDH activity, ↑TLR4, ↑CD68 cells infiltration, ↑IL-6, ↑NF-κB expression, ↑TNF-α, ↑IL-1β, caspase-3, and ↑caspase-8
LDH activity, ↓ ↓CD68 cells infiltration, ↓
↓LDH activity, ↓TLR4 content, ↓CD68 cells infiltration, ↓IL-6 content, ↓NF-κB expression,
Betulin
congestion, and
↑CRP, ↑LDH activity, ↑TLR4, ↑CD68 cells infiltration, ↑IL-6, ↑NF-kB expression, ↑TNF-α, ↑IL-1β, ↑caspase-3, and ↑caspase-8
↑ ↑LDH activity, ↑TLR4, ↑CD68 cells infiltration, ↑IL-6, ↑NF-κB expression, ↑TNF-α, ↑IL-1β, ↑ ↑
↑CRP, ↑LDH activity, ↑TLR4, ↑CD68 cells infiltration, ↑IL-6, ↑NF-κB expression, ↑TNF-α, ↑IL-1β, ↑caspase-3, and ↑caspase-8
Acetic acid-induced Sprague Dawley mice models of
Acetic acid-induced Sprague Dawley mice models of colitis
Sprague Dawley mice models of colitis
8 mg/kg/day intraperitoneally for 14
8 mg/kg/day intraperitoneally for 14 days
↓NF-κB expression, ↓TNF-α ↓IL-1β, ↓ ↓
peritoneally for 14 days
[44]
[44]
↓TNF-α expression, ↓IL-1β, ↓caspase-3 expression, and
[44]
↓TNF-α expression, ↓IL-1β, ↓caspase-3 expression, and ↓caspase-8 expression
congestion, and
caspase-8 expression
- Metabolites 2023, 13, 96 10 of 47
phosphorylation, and ↑IκBα
Table 1. Cont.
↑CD68 cells infiltration, ↑IL-6, NF-κB expression, TNF-α, ↑IL-1β, ↑caspase-3, and
8 mg/kg/day intraperitoneally for 14
NF-κB expression,
Related Molecular Mechanisms in Regulation of NF-kB in IBD
[44]
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
↓TNF-α expression, IL-1β, ↓caspase-3 expression, and ↓caspase-8 expression
mice models of colitis
submucosal edema, congestion, and immune/inflammatory
Ref.
Phytochemicals
↓TNF-α, ↓NF-kB activation, and ↑PPARγ expression in vitro and ↓pro-inflammatory cytokines TNF-α, IL-1β, and IL-6 expressions, ↓NF-kB-p65 phosphorylation, ↓IκB phosphorylation, ↑PPARγ expression, ↓phosphorylation levels of p38, ERK, and JNK, ↓NLRP3, ↓ASC, and ↓caspase-1 in vivo
NF-kB activation, and ↑PPARγ expression in vitro and ↓pro-inflammatory cytokines TNF-α, IL-1β, and IL6 expressions, NF-κB-p65 phosphorylation, ↓IκB phosphorylation, ↑PPARγ expression, phosphorylation levels of p38, ERK, and JNK,
↑TNF-α, ↑NF-kB activation, ↓PPARγ expression in vitro and ↑TNF-α, ↑IL-1β, ↑IL-6, ↑NF-kB-p65
NF-kB activation,
Naringin
Naringin
PPARγ expression in vitro and ↑TNF-α, ↑IL-1β, IL-6, ↑NF-κB-
↑Intestinal inflammation in vitro and ↑intestinal mucosa
↑Intestinal inflammation in vitro and ↑intestinal mucosa injury and ↑DAI in vivo
LPS-stimulated RAW264.7 cells in vitro and DSS-induced mice model of colitis in vivo
20 µM incubated for 1 h in vitro and 25, 50, and 100 mg/kg/day orally for 7 days in vivo
RAW264.7 cells in vitro and DSS-induced mice model of
for 1 h in vitro and 25, 50, and 100 mg/kg/day orally for 7 days
p65 phosphorylation, ↑IκB phosphorylation, ↓PPARγ
[45]
[45]
phosphorylation, ↑IκB phosphorylation, ↓PPARγ expression, ↑MAPK, ↑NLRP3, ↑ASC, and ↑caspase-1 in vivo
2023, 13, 96 9 of 44
DAI in vivo
13, 96 9 of 44
MAPK, ↑NLRP3, caspase-1 in vivo
ASC, and ↓caspase-1
ASC, and ↓caspase-1 in vivo
↓TNF-α, ↓IL-1β,
↑SOD, ↓MDA, ↓TNF-α, ↓IL-1β, ↓IL-6 and ↑IL-10 expression levels, ↓NF-kB/p65 and ↓CD3 pro-inflammatory phenotypes, ↓NF-kB/p65 mRNA, ↓MYD88, ↓p-IκBα, ↑IKKβ and ↑IκBα proteins expression, ↓NF-kB/p65 nuclear translocation
TNF-α, ↓IL-1β, ↓IL-6 and IL-10 expression levels, ↓NF-κB/p65 and ↓CD3 pro-inflammatory phenotypes, ↓NF-κB/p65 mRNA, ↓MYD88, ↓p-IκBα, ↑IKKβ and ↑IκBα proteins expression, ↓NFκB/p65 nuclear translocation
DAI,
5-Hydroxy-4-methoxycanthin-6-one
colon length, epithelial crypts destruction, disruption of the mucosal barrier, and massive submucosal
IL-10 expression levels, ↓NF-κB/p65 and ↓CD3 pro-inflammatory phenotypes, ↓NF-κB/p65 mRNA, ↓MYD88, ↓p-IκBα, ↑IKKβ and ↑IκBα proteins expression, ↓NFκB/p65 nuclear translocation
5-Hydroxy-4-methoxycanthin-6-one
Weight loss, ↑DAI, ↓colon length, epithelial crypts destruction, disruption of the mucosal barrier, and massive submucosal infiltration of inflammatory cells
↑TNF-α, ↑IL-1β, IL-6, ↓IL-10, ↓SOD, ↑MDA, NF-κB/p65, ↑CD3, ↑MYD88, p-IκBα, ↓IKKβ, ↓ NF-κB/p65 nuclear translocation
5-Hydroxy-4-methoxycanthin-6-one
↑TNF-α, ↑IL-1β, ↑IL-6, ↓IL-10, ↓SOD, ↑MDA, ↑NF-kB/p65, ↑CD3, ↑MYD88, ↑p-IκBα, ↓IKKβ, ↓IκBα, ↑NF-kB/p65 nuclear translocation
colon length, epithelial crypts destruction, disruption of the mucosal barrier, and massive submucosal
↑TNF-α, ↑IL-1β, IL-6, ↓IL-10, ↓SOD, MDA, NF-κB/p65, ↑CD3, ↑MYD88, p-IκBα, ↓IKKβ, ↓IκBα, NF-κB/p65 nuclear translocation
DSS-induced Sprague Dawley mice model of colitis
DSS-induced Sprague Dawley mice model of colitis
DSS-induced Sprague Dawley mice model of colitis
25, 50, and 100 mg/kg/day orally
25, 50, and 100 mg/kg/day orally for 11 days
mg/kg/day orally for 11 days
[46]
[46]
↑SOD, ↓IL-1β, ↓IL-6, ↓TNF-α, ↓ROS, ↑ Nrf2 activation, ↓p-NF-κBp65, and ↓p-IκBα in vitro and ↓MPO, ↑SOD, ↓IL-1β,
↓Cells viability in vitro
↓SOD, ↑IL-1β, ↑IL-6, ↑TNF-α,
↓SOD, ↑IL-1β, ↑IL-6, ↑TNF-α, ↑ROS, ↓HO-1, ↓Nrf2 activation, ↑p-NF-kBp65 and ↑p-IκBα in vitro and ↑MPO, ↓SOD, ↑IL-1β, ↑IL-6, ↑TNF-α, ↑inflammatory cells infiltration, ↓HO-1, ↓Nrf2 activation, ↑p-NF-kBp65 and ↑p-IκBα in vivo
↑SOD, ↓IL-1β, ↓IL-6, ↓TNF-α, ↓ROS, ↑HO-1, ↑Nrf2 activation, ↓p-NF-kBp65, and ↓p-IκBα in vitro and ↓MPO, ↑SOD, ↓IL-1β, ↓IL-6, ↓TNF-α, ↓inflammatory cells infiltration, ↑HO-1, ↑Nrf2 activation, ↓p-NF-kBp65 and ↓p-IκBα in vivo
↑SOD, IL-6, ↓TNF-α, ↓ROS, ↑HO-1, Nrf2 activation, ↓p-NF-κBp65, and ↓p-IκBα in vitro and ↓MPO, ↑SOD, ↓IL-1β,
IL-6, ↑TNF-α, Nrf2 activation,
Nrf2 activation, ↑p-NF-κBp65 and p-IκBα in vitro and ↑MPO, SOD, ↑IL-1β,
Geniposide
Geniposide
↓Cells viability in vitro and weight loss, ↑erosion and ↑distortion of crypts, ↑loss of glandular epithelium, and ↑inflammatory cell infiltration in vivo
↑p-NF-κBp65 and ↑p-IκBα in vitro and ↑MPO, SOD, ↑IL-1β,
RAW264.7 cells in vitro and DSS-
incubated for 24 h in vitro and 20 and 40 mg/kg/day orally/7 days in
erosion and ↑distortion of crypts, ↑loss of glandular epithelium, and
↑distortion of crypts, ↑loss of glandular epithelium, and
vitro and DSS-
in vitro and 20 and 40 mg/kg/day orally/7 days in vivo
LPS-stimulated RAW264.7 cells in vitro and DSS-induced ICR mice model of colitis in vivo
200–1000 µM incubated for 24 h in vitro and 20 and 40 mg/kg/day orally/ 7 days in vivo
[47]
↑IL-6, ↑TNF-α, inflammatory cells infiltration, HO-1, ↓Nrf2 activation, p-NF-κBp65 and
↓IL-6, ↓TNF-α, ↓inflammatory cells infiltration, ↑HO-1, ↑Nrf2 activation, p-NF-κBp65 and
induced ICR mice model of colitis in vivo
[47]
[47]
↑IL-6, ↑TNF-α, inflammatory cells infiltration, HO-1, ↓Nrf2 activation, p-NF-κBp65 and
↓IL-6, ↓TNF-α, ↓inflammatory cells infiltration, ↑HO-1, ↑Nrf2 activation, p-NF-κBp65 and
induced ICR mice model of colitis in
inflammatory cell in vivo
inflammatory cell
↑p-IκBα
↓p-IκBα in vivo
↓p-IκBα in vivo
Sesamin
↑TNF-α, ↑IL-1β, ↑IL-6, ↑p-NF- ↓TNF-α, ↓IL-1β, ↓IL-6, ↓p-NF-
infiltration
crypts destruction, levels, ↓NF-κB/p65 and ↓CD3
↓ ↓ ↑
DSS-induced 25, 50, and 100 SOD, MDA, NF-κB/p65,
Geniposide
- Metabolites 2023, 13, 96 11 of 47
↑SOD, ↓IL-1β, ↓IL-6, ↓TNF-α,
↓Cells viability in vitro
↓SOD, ↑IL-1β, ↑IL-6, ↑TNF-α,
↓p-NF-κBp65, and ↓p-IκBα in
↑erosion and
↑p-NF-κBp65 and ↑p-IκBα in
RAW264.7 cells in
incubated for 24 h
Geniposide LPS-stimulated
↓ROS, ↑HO-1, ↑Nrf2 activation,
↑ROS, ↓HO-1, ↓Nrf2 activation,
vitro and ↓MPO, ↑SOD, ↓IL-1β,
in vitro and 20 and ↑distortion of crypts, vitro and ↑MPO, ↓SOD, ↑IL-1β,
200–1000 µM and weight loss,
vitro and DSS-
vitro induced ICR mice
model of colitis vivo
Table 1. Cont.
vitro induced ICR mice model of colitis vivo
model of colitis vivo
↑IL-6, ↑TNF-α, ↑inflammatory cells infiltration, ↓Nrf2 activation, p-NF-κBp65 and
↓IL-6, ↓TNF-α, ↓inflammatory cells infiltration, HO-1, ↑Nrf2 activation, ↓p-NF-κBp65 and
↑loss of glandular epithelium, and ↑inflammatory cell
40 mg/kg/day orally/7 days in
activation, ↑p-NF-κBp65 and ↑p-IκBα in vivo
activation, ↓p-NF-κBp65 and ↓p-IκBα
↑inflammatory cell infiltration
vivo
↓ ↓ activation, p-NF-κBp65 and ↑p-IκBα in vivo
↑HO-1, ↑Nrf2 activation, p-NF-κBp65 and ↓p-IκBα in vivo
↓ Sesamin
↑
Related Molecular Mechanisms in Regulation of NF-kB in IBD
↑inflammatory cell infiltration
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
vivo
Ref.
Phytochemicals
Sesamin
↑TNF-α, ↑IL-1β, IL-6, ↑p-NFκBp65, ↑p-IκBα, ↑NF-kB
↓TNF-α, ↓IL-1β, IL-6, ↓p-NFκBp65, and ↓p-IκBα expression
↓ Sesamin
↑
p-NFκBp65, and ↓p-IκBα expression levels, ↓NF-kB signaling and activity, and MAPK levels
↓Colon length and
Sesamin
DSS-induced
50, 100, and 200
C57BL/6 mice model mg/kg/day
[48]
↓Colon length and κBp65, ↑p-IκBα, ↑NF-kB signaling and activity and ↑MAPK
Sesamin
↓IL-6, p-NFκBp65, and p-IκBα expression levels, ↓NF-kB signaling and activity, and MAPK levels
NF-kB signaling and activity, and MAPK levels
[48]
↓TNF-α, ↓IL-1β, ↓IL-6, ↓p-NF-kBp65, and ↓p-IκBα expression levels, ↓NF-kB signaling and activity, and ↓MAPK levels
IL-6, ↑p-NFκBp65, ↑p-IκBα, NF-kB signaling and activity and
↓TNF-α, ↓IL-6, ↓p-NFκBp65, and p-IκBα expression levels, ↓NF-kB signaling and activity, and MAPK levels
↑TNF-α, ↑IL-1β, ↑IL-6, ↑p-NF-kBp65, ↑p-IκBα, ↑NF-kB signaling and activity and ↑MAPK
NF-kB signaling and activity and
of colitis
orally/7 days
↓Colon length and ↓body weight
C57BL/6 mice model of colitis
mg/kg/day orally/7 days
[48]
DSS-induced C57BL/6 mice model of colitis
50, 100, and 200 mg/kg/day orally/7 days
↓Colon length and ↓body weight
↓body weight
C57BL/6 mice model of colitis
mg/kg/day orally/7 days
[48]
[48]
Taxifolin colon length,
↓TNF-α, IL-1β and ↓IL-6
Taxifolin
SIgA, ↓IL-10, ↓SOD, ↑p-NF-Kb-p65 and ↑p-IkBα
IL-1β and ↓IL-6 expression levels, ↑SIgA, ↑IL-10 and ↑SOD expression levels, ↓p-NF-Kb-p65 and ↓p-IkBα
↓body weight, ↑crypt distortion, and ↑inflammatory cells
↑TNF-α, ↑IL-1β, ↑IL-6, ↓SIgA, ↓IL-10, ↓SOD, ↑ and ↑p-IkBα
↑DAI, ↓colon length,
distortion, and
[49]
DSS-induced ICR 100 mg/kg/day
Taxifolin
IL-1β and ↓IL-6 expression levels, ↑SIgA, ↑IL-10 and ↑SOD expression levels, ↓p-NF-Kb-p65 and ↓p-IkBα
Taxifolin
↓TNF-α, ↓IL-1β and ↓IL-6 expression levels, ↑SIgA,
colon length, ↓body weight, ↑crypt distortion, and
SOD expression levels, ↓p-NF-Kb-p65 and ↓p-IkBα
[49]
Taxifolin
body weight, ↑crypt distortion, and
↑TNF-α, ↑IL-1β, IL-6, ↓SIgA, ↓IL-10, ↓SOD, p-NF-Kb-p65 and ↑p-IkBα
↓TNF-α, IL-1β and ↓IL-6 expression levels, ↑SIgA, ↑IL-10 and ↑SOD expression levels, p-NF-Kb-p65 and ↓p-IkBα
↑DAI, ↓colon length, ↓body weight, ↑crypt distortion, and ↑inflammatory cells infiltration
inflammatory cells infiltration
↑TNF-α, ↑IL-1β, ↑IL-6, ↓SIgA, ↓IL-10, ↓SOD, ↑p-NF-kB-p65 and ↑p-IkBα
↑TNF-α, ↑IL-1β, IL-6, ↓SIgA, ↓IL-10, ↓SOD, p-NF-Kb-p65
DSS-induced ICR mice model of colitis
100 mg/kg/day orally for 14 days
DSS-induced ICR mice model of colitis
100 mg/kg/day orally for 14 days
[49]
DSS-induced ICR mice model of colitis
100 mg/kg/day orally for 14 days
↑IL-10 and ↑SOD expression levels, ↓p-NF-kB-p65 and ↓p-IkBα
[49]
Body weight, ↑DAI, crypt distortion, mucosal necrosis,
inflammatory cells
inflammatory cells infiltration
Body weight, ↑DAI, ↑crypt distortion,
Isobavachalcone
↓MPO, ↓TNF-α, ↓IL-1β, ↓IL-6,
↓Body weight, ↑DAI,
↑MPO, ↑TNF-α, ↑PGE2, ↑ iNOS, ↑COX-2 and ↑p-NF-kB-p65
Isobavachalcone
↓IL-1β, ↓IL-6, ↓PGE2, ↓NO, iNOS and ↓COX-2 expression levels and ↓p-NF-kB-p65
↓Body weight, ↑DAI, ↑crypt distortion, ↑mucosal necrosis, ↑edema, ↑gland destruction, and ↑neutrophilic infiltration
Isobavachalcone
Body weight, ↑DAI, ↑crypt distortion,
PGE2, ↓ iNOS and ↓COX-2 expression levels and ↓p-NF-kB-p65
↓MPO, ↓TNF-α, ↓IL-1β, ↓IL-6, ↓PGE2, ↓NO, ↓iNOS and ↓COX-2 expression levels and ↓p-NF-kB-p65
↑MPO, ↑TNF-α, IL-1β, ↑IL-6, ↑PGE2, ↑NO, iNOS, ↑COX-2 and
mucosal necrosis, ↑edema, gland destruction, and
↑MPO, ↑TNF-α, ↑IL-1β, ↑IL-6, ↑PGE2, ↑NO, ↑iNOS, ↑COX-2 and ↑p-NF-kB-p65
↑edema, gland destruction, and
crypt distortion, ↑mucosal necrosis,
C57BL/6 mice model of colitis
mg/kg/day orally/4 days
[50]
Isobavachalcone
DSS-induced C57BL/6 mice model of colitis
25 and 50 mg/kg/day orally/4 days
↓MPO, ↓TNF-α, ↓IL-1β, ↓IL-6, ↓PGE2, ↓NO, iNOS and ↓COX-2 expression levels and ↓p-NF-kB-p65
C57BL/6 mice model of colitis
mg/kg/day orally/4 days
[50]
[50]
Isobavachalcone
↑MPO, ↑TNF-α, IL-1β, ↑IL-6, ↑PGE2, NO, iNOS, ↑COX-2 and ↑p-NF-kB-p65
↓MPO, ↓TNF-α, ↓IL-1β, ↓IL-6, ↓PGE2, ↓NO, iNOS and ↓COX-2 expression levels and ↓p-NF-kB-p65
DSS-induced
25 and 50 mg/kg/day orally/4 days
↑MPO, ↑TNF-α, IL-1β, ↑IL-6, ↑PGE2, ↑NO, and ↑p-NF-kB-p65
DSS-induced 25 and 50
mucosal necrosis, gland destruction, and
gland destruction, and
[50]
, 96 10 of 44
[50]
of colitis
infiltration
↓Body weight, ↑DAI, ↑ulcer formation, ↑thickened bowel wall, ↑hyperemia, ↑edema, and ↑mucosal inflammatory cells infiltration
↑MPO, ↑MDA, ↓GSH, ↓SOD, ↓CAT, ↑iNOS, ↑COX-2, ↑TNF-α, ↑IFN-γ, ↑IL-6, ↑IL-17, ↑IL-1β, ↓IL-10, ↓PPAR-γ and ↑NF-kB signaling
↓MPO, ↓MDA, ↑GSH, ↑SOD, ↑CAT, ↓iNOS, ↓COX-2, ↓TNF-α, ↓IFN-γ, ↓IL-6, ↓IL-17, ↓IL-1β, ↑IL-10, ↑PPAR-γ and ↓NF-kB signaling
Body weight, ↑DAI, ↑ulcer formation, ↑thickened bowel wall,
d-pinitol
d-pinitol
↓MPO, ↑GSH, ↑SOD, ↑CAT, ↓iNOS, COX-2, ↓TNFα, ↓IFN-γ, IL-6, ↓IL-17, ↓IL-1β,
↑MPO, ↑MDA, GSH, ↓SOD, ↓CAT, ↑iNOS, COX-2, ↑TNFα, ↑IFN-γ, ↑ IL-17, ↑IL-1β,
Body weight, ↑DAI, ↑ulcer formation,
d-pinitol
infiltration
DSS-induced BALB/c mice model of colitis
↓MPO, ↓MDA, GSH, ↑SOD, ↑CAT, ↓COX-2, ↓TNFα, ↓IFN-γ, IL-6, ↓IL-17, ↓IL-1β,
↑MPO, ↑MDA, GSH, ↓SOD, ↓CAT, ↑iNOS, COX-2, ↑TNFα, ↑IFN-γ, IL-17, ↑IL-1β,
10, 20, and 40 mg/kg/day
DSS-induced BALB/c mice [51] model of colitis
10, 20, and 40 mg/kg/day orally/7 days
infiltration
mg/kg/day orally/7 days
[51]
and ↑mucosal inflammatory cells
↓IL-10, ↓PPAR-γ and ↑NF-kB signaling
↑IL-10, ↑PPAR-γ and ↓NF-kB signaling
DSS-induced BALB/c mice model of colitis
↓Body weight, ↑DAI, ulcer formation, thickened bowel wall,
d-pinitol
[51]
↓MPO, ↑GSH, ↑SOD, ↑CAT, COX-2, ↓TNFα, ↓IFN-γ, ↓IL-6, ↓IL-17, ↓IL-1β,
↑MPO, ↑MDA, GSH, ↓SOD, ↓CAT, COX-2, ↑TNFα, ↑IFN-γ, ↑IL-6, ↑IL-17, ↑IL-1β,
Body weight, ↑DAI, ulcer formation,
10, 20, and 40
d-pinitol
thickened bowel wall, ↑edema,
↓MPO, ↑GSH, ↑SOD, ↑CAT, COX-2, ↓TNF-
↑MPO, GSH, ↓SOD, COX-2, ↑TNF-
↑edema,
[51]
↑thickened bowel wall,
orally/7 days
mice model of colitis
Paeoniflorin-6’-O-benzene sulfonate
Paeoniflorin-6’-O-benzene sulfonate
GRK2 translocation and ↓TLR4-NF-κB-NLRP3 inflammasome signaling in macrophages
↓GRK2 translocation and ↓TLR4-NF-kB-NLRP3 inflammasome signaling in macrophages
↑GRK2 activation and ↑TLR4NF-kB-NLRP3 inflammasome signaling
↑M1 macrophage polarization and ↑intestinal barrier dysfunction
↑GRK2 activation and ↑TLR4-NF-kB-NLRP3 inflammasome signaling
DSS-induced mice model of colitis
polarization and ↑intestinal barrier
DSS-induced mice model of colitis
17.5, 35, and 70mg/kg/day orally/6 days
[52]
70 mg/kg/day orally/6 days
[52]
- Metabolites 2023, 13, 96 12 of 47
17.5, 35, and ↑GRK2 activation and ↑TLR4-
orally/6 days signaling
↓
Table 1. Cont.
model of colitis
orally/6 days
signaling
dysfunction
macrophages
Related Molecular Mechanisms in Regulation of NF-kB in IBD
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
Ref.
Phytochemicals
AcOH-induced
10, 30, and 100
↑Intestinal inflammation and ↑OS
↑MPO, ↑TNF-α, and ↑p-NF-kBp65
↓MPO, ↓TNF-α, and ↓p-NF-kBp65
AcOH-induced
10, 30, and 100
of colitis
orally/6 days
Thymol
Thymol
↑Intestinal inflammation and ↑OS
↑MPO, ↑TNF-α, and ↑p-NF-kB- ↓MPO, TNF-α, and ↓p-NF-kB-
of colitis
orally/6 days
AcOH-induced Wistar mice model of colitis
10, 30, and 100 mg/kg/day orally/6 days
↑Intestinal inflammation and ↑OS
↑MPO, ↑TNF-α, and ↑p-NF-kB-p65
↓MPO, ↓TNF-α, and ↓p-NF-kB-p65
↑ DAI, ↓Body weight, colon length,
[53]
Wistar mice model of colitis
mg/kg/day orally/6 days
[53]
↓colon length, Inflammatory cells
infiltration, ↑epithelial cell disorganization,
12.5, 25, and 50
↓IL-6 expression, ↓TNF-α
↑ ↓Body weight, colon length, Inflammatory cells
↑ DAI, ↓Body weight, ↓colon length, ↑Inflammatory cells infiltration, ↑epithelial cell disorganization, ↑mucosal thickening, ↓crypts, ↑spleen weight, and ↑myeloid-derived suppressor cells (MDSC, CD11b+Gr1+), ↑MPO, ↑IL-6, TNF-α, and IL-1β in colonic tissues in vivo
↑NO, ↑IL-6, ↑MIP-2, ↑phosphorylated NFKb-p65 in vitro
epithelial cell disorganization, ↑mucosal thickening, ↓crypts, spleen weight, and ↑myeloid-derived suppressor cells (MDSC, CD11b+Gr1+),
mucosal thickening,
↓TNF-α
IL-1β expression Phosphorylated nuclear p65 in
↓IL-6 expression, ↓TNF-α expression, ↓MIP-2 expression, ↓IL-1β expression ↓Phosphorylated nuclear p65 in vitro and ↓NF-kB pathway in vivo
Tricin
and 100 and 150 mg/kg/day orally/7 days in vivo
[54]
spleen weight, and ↑myeloid-derived suppressor cells (MDSC, CD11b+Gr1+),
Tricin
MIP-2 expression, ↓IL-1β expression ↓Phosphorylated nuclear p65 vitro NF-κB pathway in
12.5, 25, and 50 µM incubated/30 min or 24 h in vitro and 100 and 150 mg/kg/day orally/ 7 days in vivo
12.5, 25, and 50 µM incubated/30 min or 24 h in vitro and 100 and 150 mg/kg/day
infiltration, epithelial cell disorganization, ↑mucosal thickening, ↓crypts, spleen weight,
↑NO, ↑IL-6, ↑TNF-α, ↑IL-1β, ↑MIP-2, ↑phosphorylated NFKb-p65
DSS-induced BALB/c mice model of colitis and LPS-induced RAW 264.7 treated cells
↑NO, ↑IL-6, ↑TNF-α, ↑IL-1β, ↑MIP-2, ↑phosphorylated NF-kB-p65 in vitro
IL-6 expression, ↓TNF-α expression, MIP-2 expression, ↓IL-1β expression ↓Phosphorylated nuclear p65 in
mice model of colitis and LPS-induced RAW 264.7 treated cells
min or 24 h in vitro and 100 and 150 mg/kg/day orally/7 days in
RAW 264.7 treated cells
Tricin DSS-induced BALB/c mice model of colitis and LPS-induced
vitro NF-κB pathway in vivo
[54]
↑NO, IL-6, TNF-α, ↑IL-1β, ↑MIP-2, phosphorylated NFKb-p65 in vitro
[54]
IL-6, TNF-α, and IL-1β in colonic
myeloid-derived suppressor cells (MDSC, CD11b Gr1+), ↑MPO, ↑IL-6, TNF-α, and IL-1β in colonic tissues in vivo
NF-κB pathway in
IL-6, TNF-α, and IL-1β in colonic
↓Body weight, ↑DAI, ↑colon length, ↑colon weight, inflammatory cells infiltration (mononuclear macrophages and neutrophils), ↑mucosal and submucosal lesion,
↓Body weight, ↑DAI, ↑colon length, ↑colon weight, ↑inflammatory cells infiltration (mononuclear macrophages and neutrophils), ↑mucosal and submucosal lesion, ↑degeneration, and ↑crypt cells necrosis
↑p-p65, ↑IκBα phosphorylation and ↓PPAR-γ in vitro and ↑iNOS mRNA, ↑TNF-α mRNA, ↑IL-1β mRNA, ↑p-P65, ↑MAPKs protein and phosphorylation in vivo
↑p-p65, ↑IκBα phosphorylation and ↓PPAR-γ in vitro and ↑iNOS mRNA, TNF-α mRNA, ↑IL-1β mRNA, p-P65, ↑MAPKs protein and phosphorylation in vivo
↓ ↓IL-1β mRNA, ↓p-P65, IκBα phosphorylation, ↑PPAR-γ, NK-kB activation in vitro and iNOS mRNA, ↓TNFα mRNA, IL-1β mRNA, ↓NKkB activation in vivo
200, 300, 400, and 500 µM incubated for 1 h in vitro and 1 and 5 mg/kg/day intraperitoneally every two days after colitis induction for 12 days in vivo
↓TNF-α mRNA, ↓IL-1β mRNA, ↓p-P65, ↓IκBα phosphorylation, ↑PPAR-γ, ↓NK-kB activation in vitro and ↓iNOS mRNA, ↓TNF-α mRNA, ↓IL-1β mRNA, ↓NK-kB activation in vivo
200, 300, 400, and 500 µM incubated for 1 h in vitro and 1 and 5 mg/kg/day intraperitoneally every two days after colitis induc-
for 1 h in vitro and 1 and 5 mg/kg/day intraperitoneally every two days after colitis induction for 12 days vivo
Aesculin
↓Body weight, ↑DAI, ↑colon length, ↑colon weight, inflammatory cells infiltration (mononuclear macrophages and neutrophils), ↑mucosal
Aesculin
DSS-induced BALB/c mice model of colitis and LPS-induced RAW 264.7 treated cells
mice model of colitis and LPS-induced RAW 264.7 treated cells
↑p-p65, ↑IκBα phosphorylation and ↓PPAR-γ in vitro and ↑iNOS mRNA, TNF-α mRNA, ↑IL-1β mRNA, p-P65, ↑MAPKs protein and phosphorylation in vivo
↓TNF-α mRNA, ↓IL-1β mRNA, ↓p-P65, ↓IκBα phosphorylation, ↑PPAR-γ, NK-kB activation in vitro and iNOS mRNA, ↓TNF-
200, 300, 400, and 500 µM incubated for 1 h in vitro and 1 and 5 mg/kg/day
↓Body weight, ↑DAI, ↑colon length, ↑colon weight, inflammatory cells infiltration (mononuclear
Aesculin DSS-induced BALB/c mice model of colitis and LPS-induced
[55]
[55]
↑p-p65, ↑IκBα phosphorylation and ↓PPAR-γ in vitro and ↑iNOS mRNA, TNF-α mRNA, ↑IL-1β mRNA, ↑p-P65,
↓TNF-α mRNA, ↓IL-1β mRNA, ↓p-P65, ↓IκBα phosphorylation, ↑ NK-kB activation in vitro and ↓iNOS mRNA, ↓TNF-
Aesculin DSS-induced BALB/c
[55]
↓ kB activation in vivo
cells
RAW 264.7 treated
every two days af-
- Metabolites 2023, 13, 96 13 of 47
Table 1. Cont.
Related Molecular Mechanisms in Regulation of NF-kB in IBD
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
Ref.
Phytochemicals
↑degeneration, and
↑
↑
↑ ↑claudin occludin mRNA,
↑ ↑claudin
↑ZO-1 mRNA, ↑claudin mRNA, ↑occludin mRNA, ↓TNF-α mRNA, ↓IL-1β mRNA, ↓IL-6 mRNA, ↓MCP-1 mRNA, ↓F4/80
↑claudin
Ginsenoside Rk3
Ginsenoside Rk3
Body weight, ↑ ↑
↓IL-1β mRNA, ↓IL-6 mRNA, MCP-1 mRNA, ↓F4/80 mRNA, ↓NADPH mRNA, ↓STAMP2 mRNA,
TNF-α mRNA, ↓IL-1β mRNA, ↓IL-6 mRNA, ↓ ↓ mRNA, ↓
TNF-α mRNA, ↓IL-1β mRNA, ↓IL-6 mRNA, ↓ ↓ mRNA, ↓
↑Body weight, ↑fat accumulation, ↑glucose tolerance, ↓colon length, ↑inflammatory cells infiltration and ↑crypt lesions
accumulation, ↑glucose tolerance, colon length, inflammatory cells infiltration,n and
↓ZO-1 mRNA, mRNA, ↓occludin mRNA,↑TLR4, MYD88, and ↓IkBα
↑ colon length, inflammatory
↓ZO-1 mRNA, mRNA, ↓ mRNA,↑TLR4, ↓IkBα
↓ZO-1 mRNA, claudin mRNA, ↓ mRNA,↑ ↓IkBα
HFD-induced obese C57BL/6 mice model of colitis
↓ZO-1 mRNA, ↓claudin mRNA, ↓occludin mRNA,↑TLR4, ↑MYD88, and ↓IkBα
HFD-induced obese C57BL/6 mice model of colitis
colon inflammatory
30 and 60 mg/kg/day orally/8 weeks
[56]
mg/kg/day orally/8 weeks
[56]
mRNA, ↓NADPH mRNA, ↓STAMP2 mRNA, ↓TLR4, ↓JNK/phosphorylation
JNK/phosphorylation TLRA4/MYD88 IkBα mRNA
JNK, ↓NF-kB, ↓TLRA4/MYD88 pathway, and ↑IkBα mRNA
IkBα
IkBα
Lancemaside A
Lancemaside A
↓TLR4-linked NF-kB in vitro and ↓TNF-α mRNA, ↓IL-1β mRNA, ↓IL-6 mRNA, TLR4 mRNA, ↓NF-κB-p65 mRNA and ↓COX2 mRNA
↓TLR4-linked NF-kB in vitro and ↓TNF-α mRNA, ↓IL-1β mRNA, ↓IL-6 mRNA, TLR4 mRNA, ↓NF-κ ↓
↓TLR4-linked NF-kB in vitro and ↓TNF-α mRNA, ↓IL-1β mRNA, ↓IL-6 mRNA, TLR4 mRNA, ↓NF-κ ↓
↑TLR4-linked NF-kB in vitro and ↑MPO, ↑TNF-α mRNA and ↑IL-1β mRNA, ↑IL-6 mRNA, ↑TLR4 mRNA, ↑NF-kB (pp65) mRNA, ↑COX-2 mRNA in vivo
↓TLR4-linked NF-kB in vitro and ↓TNF-α mRNA, ↓IL-1β mRNA, ↓IL-6 mRNA,
TLR4-linked NF-kB in vitro and ↑MPO, ↑TNF-α mRNA and ↑IL-1β mRNA, IL-6 mRNA, ↑TLR4 mRNA, NF-κB (pp65) mRNA, COX-2 mRNA in vivo
TLR4-linked NF-kB in vitro and ↑ ↑TNF-α mRNA and ↑IL-1β mRNA, IL-6 mRNA, ↑TLR4 mRNA, NF-κ mRNA, in vivo
↑TLR4-linked NF-kB in vitro and ↑ ↑TNF-α mRNA and ↑IL-1β mRNA, IL-6 mRNA, ↑TLR4 mRNA, NF-κ mRNA, in vivo
mice model of colitis and LPS-induced 293-hTLR4Ahemagglutinin treated cells
mice model of colitis and LPS-induced in vitro
↓Colon length, ↑thicken, ↑erythematous colon, ↑edema, ↑inflammatory cells infiltration, and ↑epithelial ulcers
TNBS-induced ICR mice model of colitis and LPS-induced 293-hTLR4Ahemagglutinin treated cells
↑ ↑ ↑edema, inflammatory
mice model of colitis and LPS-induced in vitro
20 µM and 100µM incubated for 6 h in vitro and 10 or 20 mg/kg/day orally for 5 days in vivo
↑erythematous colon, ↑edema, inflammatory cells infiltration, and
↑ ↑edema, inflammatory
in vitro and 10 or 20 mg/kg/day orally for 5 days in
[57]
[57]
↓TLR4 mRNA, ↓NF-kB-p65 mRNA and ↓COX-2 mRNA in vivo
in
in
epithelial ulcers
↓NF-κB translocation into the nucleus, ↑NF-κB P65 protein in the cytoplasm, ↓nuclear NF-kB P65 protein levels, ↓INF-γ expression, ↑P65 in the cytoplasm, P65 in the nucleus in vitro and ↓NF-kB P65 in the nucleus, ↓p-IKBα, ↑NF-kB in the cytoplasm, ↓nuclear NF-κB p65 protein levels, ↓C-MYC expression, iNOS expression
↓NF-κB translocation into the nucleus, ↑NF-κ the cytoplasm, ↓
↓NF-kB translocation into
NF-κB translocation into the nucleus, ↑NF-κ the cytoplasm, ↓
NF-κB translocation into the
NF-κB translocation into the
↑NF-kB translocation into
the nucleus, ↑NF-kB P65 protein in the cytoplasm, ↓nuclear NF-kB P65 protein
NF-κB translocation into the nucleus, ↓
the nucleus, ↓NF-kB P65 protein in the cytoplasm, ↑nuclear NF-kB P65 protein
P65 protein in the cytoplasm, ↑nuclear NFkB P65 protein levels, ↑TNF- α, ↑IL-6, ↑IL-8, INF-γ mRNA and ↑ROS ↑NF-kB P65 in the nucleus, ↓NF-kB in cytoplasmic, C-MYC expression, iNOS expression,
diarrhea with or without hematochezia, ↑DAI, ↑inflammation of
↓Body weight, ↑diarrhea with or without hematochezia, ↑DAI, ↑inflammation of the mucosa, ↑fibrotic thickening, ↑ulcers, ↑edema, ↑microhemorrhages, and ↑ necrosis
↑nuclear NF-
↓INF-γ expression, ↑
Tetramethylpyrazine
Tetramethylpyrazine
↑nuclear NF-
↓INF-γ expression, ↑
levels, ↓INF-γ expression, ↑P65 in the cytoplasm, ↓P65 in the nucleus in vitro and ↓NF-kB P65 in the nucleus, ↓p-IKBα, ↑NF-kB in the cytoplasm, ↓nuclear NF-kB p65 protein levels, ↓C-MYC expression, ↓iNOS expression and ↓COX-2 expression in vivo
↑ α, ↑ ↑ INF-γ mRNA and ↑ ↑
40 µg/mL incubated for 24 h in vitro and 80 mg/kg/day intraperitoneally/7 days in vivo
bated for 24 h in vitro and 80 mg/kg/day intraperitoneally/7
in vitro
↑ α, ↑ ↑ INF-γ mRNA and ↑ ↑
in vitro
levels, ↑TNF- α, ↑IL-6, ↑IL-8, ↑INF-γ mRNA and ↑ROS in vitro, and ↑NF-kB P65 in the nucleus, ↓NF-kB in cytoplasmic, ↑C-MYC expression, ↑iNOS expression, ↑COX-2 expression in vivo
↑DAI,
Oxazolone-induced KM mice model of colitis and LPS-treated Caco-2 cells
KM mice model of colitis and LPStreated Caco-2 cells
↑ thickening,
[58]
fibrotic thickening, ulcers,
[58]
↑ thickening,
in vitro and nucleus, ↓ α, ↑NF-kB in the cytoplasm, ↓nuclear NF-κ
in vitro and nucleus, ↓ α, ↑NF-kB in the cytoplasm, ↓nuclear NF-κ
↓NF-kB in C-MYC
↓NF-kB in C-MYC
microhemorrhages,
↓C-MYC expression,
expression, COX-2 expression in vivo
↓C-MYC
COX-2 expression in vivo
expression, and COX-2 expression in vivo
COX-2 expression in vivo
COX-2 expression in vivo
COX-2 expression in vivo ↑ ↑
↑DAI, ↑diarrhea,
hemagglutinin orally for 5 days in cells infiltration, and NF-κB-p65 mRNA and COX-
- Metabolites 2023, 13, 96 14 of 47
in the cytoplasm, ↑nuclear NF-
P65 protein levels, ↓INF-γ
40 µg/mL incu-
Tetramethylpyrazine
without hematochezia,
Table 1. Cont.
colitis and LPStreated Caco-2 cells
thickening, ulcers,
nucleus, ↓p-IKBα, ↑NF-kB in the cytoplasm, nuclear NF-κB p65 protein levels, ↓C-MYC
in the nucleus, ↓NF-kB in cytoplasmic, expression, iNOS expression,
peritoneally/7 in vivo
Related Molecular Mechanisms in Regulation of NF-kB in IBD
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
↑microhemorrhages, and ↑
Ref.
Phytochemicals
iNOS expression and ↓COX-2 expression in vivo
↑NO, ↑ROS, ↓GSH, ↑NF-kB-p65, ↑p65, ↓IkBα in vitro and ↑NO, ↑ COX-2, ↑PGE2, ↑IL-1β, ↑MMP-9, ↓IL-4, ↓IL-10, Gene expression of ↑Wnt-1, ↑β-Catenin, ↑cyclin-D1, ↑C-MYC, ↑Expression of Wnt-1, β-Catenin and LRP6, ↓Expression of p-GSK3β and ↑Expression of NF-kB p65 and p-IkBα in vivo
↓NF-kB p65, ↓p65, ↑IkBα in vitro and Gene expression of ↓Wnt-1, ↓β-Catenin, ↓cyclin-D1, ↓C-MYC, ↓GSK3β, ↑Expression of TCF-4, LEF-1 and p-GSK3β, ↓Expression of Wnt-1, β-Catenin and LRP6 and ↓expression of NF-kB p65 and p-IkBα in vivo
↑NO, ↑ROS, ↓GSH, ↑NF-kBp65, ↑p65, ↓IkBα in vitro and ↑NO, ↑ COX-2, PGE2, ↑IL-1β, ↑MMP-9, IL-10, Gene expression of Wnt-1, ↑βCatenin, ↑cyclin-D1, ↑C-MYC, ↑Expression of Wnt-1, βCatenin and LRP6, ↓Expression
↓NF-kB p65, p65, ↑IkBα in vitro and Gene expression of ↓Wnt-1, ↓βCatenin, ↓cyclin-D1, ↓C-MYC, ↓GSK3β, ↑Expression of TCF-4, LEF-1 and p-GSK3β, Expression of Wnt-1, β-Catenin and LRP6
↑DAI, ↑diarrhea, ↑bleeding, ↓colon length, ↑edema, ↑congestion, ↑thickening, ↑erosion, ↑ulceration, ↑adhesions to adjacent tissues, ↑mucosal damage, ↑inflammatory cell infiltration, ↑crypt loss, and ↑TUNEL stained spots
colon length, edema, ↑congestion, ↑thickening, ↑erosion, ↑ulceration, adhesions
Daurisoline
Daurisoline
20, 50, and 100 µM incubated for 24 h in vitro and 10, 20, 40 mg/kg/day orally/7 days in
0, 0.5, 1, 2, 5, 10, 20, 50, and 100 µM incubated for 24 h in vitro and 10, 20, 40 mg/kg/day orally/7 days in vivo
DSS-induced BALB/c mice model of colitis and LPS-induced RAW 264.7 treated cells
12 of 44
mice model of colitis and LPS-induced RAW 264.7 treated
12 of 44
[59]
[59]
to adjacent tissues, mucosal damage, inflammatory cell
and ↓expression of NF-kB p65 and p-IkBα in vivo
crypt loss, TUNEL stained
of p-GSK3β and Expression of NF-kB p65 and p-IkBα in vivo
and ↓expression of NF-kB p65 and p-IkBα in vivo
crypt loss, TUNEL stained
of p-GSK3β and Expression of NF-kB p65 and p-IkBα in vivo
Tetrandrine
Tetrandrine
Tetrandrine
↓NF-kB DNA bindng activity, ↓IL-1β mRNA and protein, ↓TNF-α mRNA and protein
NF-kB DNA binding activity, IL-1β mRNA and protein, TNF-α mRNA and protein,
↓NF-kB DNA bindng activity, ↓IL-1β mRNA and protein, ↓TNF-α mRNA and protein
↑NF-kB DNA binding activity, ↑IL-1β mRNA and protein, ↑TNF-α mRNA and protein, and ↑MPO
IL-1β mRNA and protein, TNF-α mRNA and protein,
↓NF-kB DNA bindng activity, ↓IL-1β mRNA and protein, ↓TNF-α mRNA and protein
↑DAI
[60]
DSS-induced BALB/c mice model of colitis
40 mg/kg/day orally/7 or 14 days
DSS-induced BALB/c mice model of colitis
40 mg/kg/day orally/7 or 14 days ↑DAI
mice model of colitis orally/7 or 14 days
↑DAI
[60]
[60]
↑DAI, ↓body weight, colonic damage,
↑ ↓body weight, colonic damage,
↑DAI, ↓body weight, ↑colonic damage, ↑ulceration, ↑stool consistency score, ↑destruction of colon tissue, ↑inflammatory cell infiltration, ↑necrosis and ↑edema
↓GSH, ↓SOD, ↑MDA, ↑NO, ↑MPO, ↑hydroxyproline, ↑TNF-α, ↑IL-1β, ↑IL-6, ↓IL-10, ↑iNOs mRNA, ↑IFN-γ mRNA, ↑COX-2 mRNA, ↑LTB4 mRNA, ↑Bax, ↑Caspase-1, ↑NF-kB and ↑IκBα,
↓GSH, ↓SOD, MDA, ↑NO, ↑MPO, hydroxyproline, ↑TNFα, ↑IL-1β, IL-10, ↑iNOs mRNA, ↑IFN-γ mRNA, ↑COX2 mRNA, LTB4 mRNA, ↑Bax, ↑Caspase-1, ↑NF-kB and ↑IκBα,
Diosgenin
Diosgenin
↓GSH, MDA, ↑NO, ↑MPO, hydroxyproline, ↑TNFα, ↑IL-1β, ↑ IL-10, ↑iNOs mRNA, ↑IFN-γ mRNA, ↑COX2 mRNA, ↑LTB4 mRNA, ↑Bax,
↑stool consistency score, ↑destruction of colon
Diosgenin
↓iNOs mRNA, ↓COX-2 mRNA, ↓IFN-γ mRNA, ↓Bax, ↓Caspase-1, ↓NF-kB
↓iNOs mRNA, ↓COX-2 mRNA, ↓IFN-γ mRNA, ↓Bax, ↓Caspase-1, NF-kB and ↓IκBα
TNBS-induced Sprague-Dawley rat model of colitis
stool consistency score,
TNBS-induced Sprague-Dawley rat model of colitis
↓iNOs mRNA, ↓COX-2 mRNA, ↓IFN-γ mRNA, ↓Bax, ↓Caspase-1, NF-kB and ↓
TNBS-induced Sprague-Dawley rat
50, 100, or 200 mg/kg/day
50, 100, or 200 mg/kg/day orally/14 days
mg/kg/day orally/14 days
[61]
[61]
[61]
destruction of colon inflammatory
and ↓IκBα
inflammatory cell infiltration, ↑necrosis and ↑edema
NF-kB and ↑IκBα,
cell infiltration, ↑ ↑
↑ ↑
- Metabolites 2023, 13, 96 15 of 47
↑colonic damage,
↓GSH, ↓SOD, ↑MDA, ↑NO,
Table 1. Cont.
Sprague-Dawley rat model of colitis orally/14 days
↓Caspase-1, ↓NF-kB and ↓IκBα
tissue, ↑inflammatory cell infiltration, ↑necrosis and edema
2 mRNA, ↑LTB4 mRNA, ↑Bax, ↑Caspase-1, ↑IκBα,
Related Molecular Mechanisms in Regulation of NF-kB in IBD
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
Ref.
Phytochemicals
IRAK1 phosphorylation and NF-kB
↓IRAK1 phosphorylation and degradation, ↓NF-kB activation, ↓IKKβ phosphorylation, ↓IκBα phosphorylation and degradation, ↓p65 translocation, ↓MAPK p38 phosphorylation, ↓ERK phosphorylation, ↓JNK phosphorylation, ↓TNF-α expression, ↓IL-1β expression, ↓IL-6 expression, ↓COX-2 expression, ↓iNOS expression and ↑IL-10 expression in vitro and ↓phosphorylation of IRAK1 and IKKβ, ↓NF-kB activation, ↓TNF-α expression, ↓IL-1β expression and ↓IL-6 expression in vivo
IKKβ phosphorylation, ↓IκBα phosphorylation and degradation, p65 translocation, MAPK p38 phosphorylation, ↓ERK phosphorylation, ↓JNK phosphorylation, ↓TNF-α expression, IL-1β expression, ↓IL-6 expression, ↓COX-2
↑IRAK1 phosphorylation and degradation, ↑degradation of IRAK1, 2, and 4, ↑NF-kB activation, ↑TAK1 phosphorylation and degradation, ↑IKKβ phosphorylation, ↑IκBα phosphorylation and degradation, ↑PGE2, ↑NO, ↑TNF-α expression, ↑IL-1β expression, ↑IL-6 expression, ↑IL-10 expression, ↑COX-2, ↑iNOS expression in vitro and ↑IRAK1 phosphorylation in vivo
degradation of NF-kB
phosphorylation and IKKβ
Mangiferin
Mangiferin
5, 10, and 20 µM incubated for 15 to 120 min in vitro and 10 or 20 mg/kg/day orally/3 days in vivo
TNBS-induced C57BL/6 mice model of colitis and LPS-induced peritoneal macrophages
↑IκBα phosphorylation and
C57BL/6 mice model of colitis and LPSinduced peritoneal macrophages
120 min in vitro and 10 or 20 mg/kg/day orally/3 days in
↓Colon length, ↑MPO
[62]
↓Colon length, ↑MPO
PGE2, ↑NO, TNF-α expression, ↑IL-1β
expression, iNOS expression and ↑IL-10 expression in vitro and
IL-6 expression, ↑COX-2, in vitro and
↓phosphorylation of IRAK1
IRAK1 phosphorylation in
13 of 44
NF-κB activation, TNF-α expression, ↓IL-1β
↓IL-6 expression in vivo
Tryptanthrin
↑TNF-α, ↑IL-1β, IL-6, ↓IL-10, ↑NF-kBp65, ↑p-STAT3, ↓IκBα protein, STAT3 and ↑p-STAT3
crypts and goblet cells, erosive lesions, ↑inflammatory
cell infiltration, and ↑atrophy
↑CAS, ↓crypts and goblet cells, ↑erosive lesions, ↑inflammatory cell infiltration, and ↑atrophy
↑TNF-α, ↑IL-1β, ↑IL-6, ↓IL-10, ↑NF-kBp65, ↑p-STAT3, ↓IκBα protein, ↑STAT3 and ↑p-STAT3
↓NF-kBp65, ↓IκBα degradation
Tryptanthrin
156.8 mg/kg twice a day orally/8 days
[63]
DSS-induced mice model of colitis
39.2, 78.4, and 156.8 mg/kg twice a day orally/8 days
↓NF-kBp65, ↓p-STAT3, and ↓IκBα degradation
model of colitis
[63]
↑ ↑ ↑IL-6, iNOS mRNA,
↑TNF-α, ↑IL-1β, ↑IL-6, ↑COX2 mRNA, ↑iNOS mRNA, ↓Ki67-positive cells,
iNOS mRNA,
Ki67-positive cells, ↑TUNELpositive cells, Occludin mRNA, ↓Claudin1 mRNA, ↓Ecadherin mRNA, ↑ p65, ↑pp65, ↑p53,
↓COX2 mRNA, iNOS mRNA, ↑Occludin mRNA, ↑Claudin1 mRNA, ↑Ecadherin mRNA, ↓p65, ↓p-p65, ↓p53, p-p53, and ↓p-AKT expression
↓COX2 mRNA, ↓iNOS mRNA, ↑Occludin mRNA, ↑Claudin1 mRNA, ↑Ecadherin mRNA, ↓p65, ↓p-p65, ↓p53, ↓p-p53, and ↓p-AKT expression
l-Theanine
Ki67-positive cells, ↑TUNELpositive cells, ↓Occludin mRNA, ↓Claudin1 mRNA, ↓Ecadherin mRNA, ↑ p65, ↑p-
↓COX2 mRNA, ↓iNOS mRNA, ↑Occludin mRNA, ↑Claudin1 mRNA, ↑Ecadherin mRNA, ↓p65, ↓p53, p-p53, and ↓p-AKT expression
l-Theanine
Body weight, ↓length of colon, colon weight, ↑DAI, ↑inflammatory infiltrates, and
↓Body weight, ↓length of colon, ↓colon weight, ↑DAI, ↑inflammatory infiltrates, and ↑epithelial injury
↑TUNEL-positive cells, ↓Occludin mRNA, ↓Claudin1 mRNA, ↓Ecadherin mRNA, ↑ p65, ↑p-p65, ↑p53, ↑p-p53 and ↑p-AKT expression and ↑lipid metabolic perturbation
colon weight, ↑DAI, ↑inflammatory infiltrates, and
l-Theanine
DSS-induced C57BL/6J mice model of colitis
Water contained 0.1% of l-theanine for 14 days orally
DSS-induced C57BL/6J mice
Water contained 0.1% of l-theanine
C57BL/6J mice model of colitis
0.1% of l-theanine for 14 days orally
[64]
epithelial injury
p-AKT expression lipid metabolic
p-AKT expression and ↑lipid metabolic
perturbation
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
Mechanisms in Regulation of NF-kB in IBD
Ref.
Phytochemicals
↓iNOS expression and ↓COX-2 expression, ↓IL-6 mRNA, ↓TNF-α mRNA, ↓MyD88-dependent TLR4 pathway, ↓DNA binding of AP-1, ↓DNA-binding activity of NF-kB, ↓c-Fos phosphorylation, ↓phosphorylation and nuclear translocation of p65. ↓phosphorylation and
↑NO, ↑PGE2, ↑expression of iNOS and ↑ expression of COX-2, ↑IL-6 mRNA, ↑TNF-α mRNA, ↑AP-1, ↑DNA-binding activity of NF-kB in vitro and ↑F4/80 mRNA, ↑Ly6G mRNA, ↓ZO-1 mRNA, ↓occludin mRNA, ↑claudin-1 mRNA, ↓E-cadherin mRNA, ↑N-cadherin mRNA, ↑vimentin mRNA, ↑iNOS mRNA, ↑COX-2 mRNA, ↑IL-6 mRNA, ↑TNF-α mRNA, ↑c-Fos, ↑p65, STAT1 and ↑STAT3 phosphorylation in vivo
degradation of IκBα, ↓phosphorylation of
↑DAI, ↑body weight loss, ↑stool consistency, ↑occult fecal blood, ↓colon length, ↑spleen index, ↑mucosal layer, ↑ulceration, ↑crypt loss, and ↑inflammatory cell infiltration
IKKα/β, ↓phosphorylation of TAK1, ↓STAT1 (Y701 and S727), ↓STAT3 (Y705),
LPS-induced RAW 264.7 and peritoneal macrophages treated cells and DSS-induced ICR mice model of colitis
20, 40, or 80 µM in vitro incubated/4 days and 5 or 20 mg/kg/day orally/ 7 days in vivo
[65]
Koreanaside A
↓JAK1 (Y1022), JAK2 (Y1007/1008) phosphorylation in vitro and ↓F4/80 mRNA, ↓Ly6G mRNA, ↑ZO-1 mRNA, ↑occludin mRNA, ↓claudin-1 mRNA, ↑E-cadherin mRNA, ↓N-cadherin, ↓iNOS mRNA, ↓COX-2 mRNA, ↓IL-6 mRNA, ↓TNF-α mRNA, ↓vimentin mRNA, ↓↑c-Fos, p65, STAT1 and ↑STAT3 phosphorylation in vivo
14 of 44
↓Body weight, ↓crypt
6-gingerol
6-gingerol
↓Body weight, ↓crypt cells, ↓goblet, ↑granulation, ↑hyperplasia, and ↑inflammatory cells infiltration
↑granulation, hyperplasia, and inflammatory cells
↑IL-17, ↓IL-10, IkBα, ↑p65, ↑pIκBα and
↓IkBα, ↓p65, p-IκBα and ↓pp65
DSS-induced BALB/c mice model of colitis
DSS-induced BALB/c mice [66] model of colitis
100 and 250 mg/kg/day orally/14 days
↑IL-17, ↓IL-10, ↑IkBα, ↑p65, ↑p-IκBα and ↑p-p65
↓IkBα, ↓p65, ↓p-IκBα and ↓p-p65
mg/kg/day orally/14 days
[66]
infiltration
↓ ↓
Table 1. Cont.
DSS-induced BALB/c mice model of colitis
DSS-induced BALB/c
granulation,
IL-17, IL-10, IkBα, p65, p- IkBα, p65, p-IκBα and p-
↑hyperplasia, and
IκBα and ↑p-p65 p65
100 and 250
↑granulation,
↑IL-17, ↓IL-10, ↑IkBα, ↑p65, ↑pIκBα and ↑
↓IkBα, ↓p65, ↓p-IκBα and ↓pp65
DSS-induced BALB/c mice model of colitis
mg/kg/day orally/14 days
[66]
↑granulation, ↑hyperplasia, and ↑inflammatory cells
↑IL-17, ↓IL-10, ↑IkBα, ↑p65, ↑pIκBα and
↓IkBα, ↓p65, ↓p-IκBα and ↓pp65
orally/14 days
DSS-induced BALB/c mice model of colitis
[66]
Related Molecular Mechanisms in Regulation of NF-kB in IBD
[66]
hyperplasia, and ↑inflammatory cells
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
↑inflammatory cells infiltration
orally/14 days
infiltration
orally/14 days
Ref.
Phytochemicals
↓Body weight, DAI, ↑colon length, ↑colon
↑MPO, ↓SOD, CAT, ↓GSH-Px, ↑MDA, ↑IFN-γ, ↑TNF-α, ↑IL-6, ↑IL-1β, ↑ TRIF, and ↑pNF-κB p65 expression, ↓ZO-1, ↓occludin, and ↓claudin-1 expressions
↓Body weight, ↑DAI,
↑MPO, ↓SOD, ↓ CAT, ↓GSH-Px, ↑MDA, ↑ TNF-α, ↑IL-6, ↑IL-1β, ↑TLR4, TRIF, and ↑pNF-κB p65 expression, ↓ZO-1, ↓occludin, and ↓claudin-1
↓TLR4, ↓TRIF, ↓p-NF-κB p65 expression, and ↑claudin-1 expressions
↓Body weight, ↑DAI,
↑MPO, ↓SOD, ↓ CAT, ↓GSH-Px, ↑MDA, ↑IFN-γ, TNF-α, ↑IL-6, ↑IL-1β, ↑TLR4, TRIF, and ↑pNF-κB p65 expression, ↓ZO-1, ↓occludin, and claudin-1
Lycopene
CAT, ↓GSH-Px, ↑MDA, ↑ TNF-α, ↑IL-6, ↑IL-1β, ↑TLR4, TRIF, and ↑pNF-κB p65 expression, ↓ZO-1, ↓occludin, and claudin-1
↑MPO, ↓SOD, ↓ CAT, ↓GSH-Px, ↑MDA, ↑IFN-γ, ↑TNF-α, ↑IL-6, ↑IL-1β, ↑TLR4, ↑TRIF, and ↑p-NF-kB p65 expression, ↓ZO-1, ↓occludin, and ↓claudin-1 expressions
Lycopene
↑colon glandular
↓TLR4, ↓ expression, ZO-1, ↑occludin and ↑claudin-1 expressions
glandular disorder, and ↑inflammatory cell
Lycopene
Lycopene
colon length, ↑colon weight, glandular disorder, and
↓TLR4, ↓TRIF, p-NF-κB p65 expression, ZO-1, ↑occludin and
↓Body weight, ↑DAI, ↑colon length, ↑colon weight, ↑glandular disorder, and ↑inflammatory cell infiltration
colon length, ↑colon weight, glandular
↓TLR4, ↓TRIF, p-NF-κB p65 expression, ZO-1, ↑occludin and
[67]
↓TLR4, ↓TRIF, ↓p-NF-kB p65 expression, ↑ZO-1, ↑occludin and ↑claudin-1 expressions
DSS-induced C57BL/6 mice model of colitis
5, 10, and 20 mg/kg/day orally/14 days
C57BL/6 mice model of colitis
mg/kg/day orally/14 days
[67]
of colitis orally/14 days
DSS-induced C57BL/6 mice model of colitis
5, 10, and 20 mg/kg/day orally/14 days
C57BL/6 mice model of colitis
mg/kg/day orally/14 days
[67]
disorder, and inflammatory cell
[67]
[67]
↑claudin-1 expressions
inflammatory cell
↑claudin-1 expressions
inflammatory cell
Body weight, ↑DAI,
↓Body weight, ↑DAI,
↑diarrhea, ↑bleeding, colon length, ulceration, erosion,
α-mangostin
↓Body weight, ↑DAI,
↑MPO, ↑phosphorylation of IKKα and IκB, activated NFκB, ↑MAPK, ↑phosphorylation of ERK1/2, SAPK/JNK and p38
↓IKKα phosphorylation, ↓IκBα
↓ ↑DAI,
α-mangostin
↓IKKα phosphorylation, ↓IκBα phosphorylation, ↓activated NF-kB, ↓phosphorylation of ERK1/2, SAPK/JNK and ↓p38
↓Body weight, ↑DAI, ↑diarrhea, ↑bleeding, ↓colon length, ↑ulceration, ↑erosion, ↑crypt distortion, ↑inflammatory cell infiltration, and ↑edema
↑MPO, ↑phosphorylation of IKKα and IκB, ↑activated NF-kB, ↑MAPK,
α-mangostin
bleeding, colon length,
α-mangostin
↑MPO, ↑phosphorylation of IKKα and IκB, activated NFκB, ↑MAPK, ↑phosphorylation of ERK1/2, SAPK/JNK and p38
↓IKKα phosphorylation, ↓IκBα
bleeding, ↓colon length, ↑ erosion,
phosphorylation, ↓activated NF-κB, phosphorylation of ERK1/2, SAPK/JNK and ↓p38
α-mangostin
bleeding, ↓colon length,
↑MPO, ↑phosphorylation of IKKα and IκB, activated NFκB, ↑MAPK, ↑phosphorylation of ERK1/2, SAPK/JNK and p38
↓IKKα phosphorylation, ↓IκBα
30 and 100 mg/kg/day orally/14 days
↑MPO, ↑phosphorylation of IKKα and IκB, activated NFκB, ↑MAPK, phosphorylation of ERK1/2, SAPK/JNK and p38
↓IKKα phosphorylation, ↓IκBα
mg/kg/day orally/14 days
[68]
phosphorylation, ↓activated NF-κB, phosphorylation of ERK1/2, SAPK/JNK and ↓p38
30 and 100 mg/kg/day
DSS-induced mice model of colitis
DSS-induced mice model of colitis
30 and 100 mg/kg/day orally/14 days
model of colitis
30 and 100 mg/kg/day
phosphorylation, ↓activated NF-κB, phosphorylation of ERK1/2, SAPK/JNK and ↓
↑ulceration, erosion, crypt distortion, inflammatory cell
DSS-induced mice model of colitis
[68]
[68]
↑crypt distortion, inflammatory cell infiltration, and ↑edema
phosphorylation, ↓activated NF-κB, phosphorylation of ERK1/2, SAPK/JNK and
DSS-induced mice
↑phosphorylation of ERK1/2, SAPK/JNK and p38
[68]
erosion, crypt distortion, ↑inflammatory cell
[68]
crypt distortion, inflammatory cell
infiltration, and ↑edema
Ophiopogonin D
Ophiopogonin D
COX-2,
↑cl-caspase3 and ↑COX-2, ↑MLCK and ↑iNOS in vitro and ↑TNF-α, ↑IL-6, ↑IL-1β, ↓Bcl-2, ↓occludin, ↑NF-kB-p65, ↑cl-caspase3, ↑Bax, ↑MLCK, ↑MDA, ↓GSH, ↓SOD, ↑iNOS, ↑COX-2 in vivo
Ophiopogonin D
COX-2, ↑MLCK and ↑iNOS in vitro and ↑TNF-α, ↑IL-6, IL-1β, ↓Bcl-2, ↓occludin, ↑NF-Κb-p65, ↑clcaspase3, MLCK,
Ophiopogonin D
iNOS in vitro and ↑TNF-α, ↑IL-6, IL-1β, ↓Bcl-2, ↓occludin, ↑NF-Κb-p65, ↑clcaspase3, ↑Bax, MLCK, ↑MDA, SOD, ↑iNOS,
cl-caspase3 and COX-2, ↑MLCK and ↑iNOS in vitro and ↑TNF-α, ↑IL-6, ↑IL-1β, ↓Bcl-2, ↓occludin, ↑NF-Κb-p65, ↑cl-
Ophiopogonin D
10 mg/kg and 40 mg/kg/day orally/7 days in vivo 20 µmol/L
Ulceration, ↑congestion, ↑edema, ↑inflammatory cell
↑congestion, ↑edema, ↑inflammatory cell infiltration, colon
↑cl-caspase3 and COX-2, ↑MLCK and ↑iNOS in vitro and ↑TNF-α, ↑IL-6, IL-1β, ↓Bcl-2, ↓occludin, NF-Κb-p65, ↑cl-
mg/kg/day orally/7 days in vivo 20 µmol/L incubated for 24 h
↑Ulceration, ↑congestion, ↑edema, ↑inflammatory cell
10 mg/kg and 40 mg/kg/day orally/7 days in vivo 20 µmol/L
DSS-induced C57BL/6J mice model of colitis and LPS-induced IEC-6 treated cells
10 mg/kg and 40 mg/kg/day orally/7 days in vivo 20 µmol/L incubated for 24 h in vitro
↑Ulceration, ↑congestion, ↑edema, ↑inflammatory cell infiltration, ↓colon length, and ↓body weight
DSS-induced C57BL/6J mice model of colitis and LPS-induced IEC-6
↑Ulceration, ↑congestion, ↑edema, ↑inflammatory cell
C57BL/6J mice model of colitis and LPS-induced IEC-6
10 mg/kg and 40 mg/kg/day orally/7 days in
DSS-induced C57BL/6J mice model of colitis and
DSS-induced C57BL/6J mice model of colitis and
↓NF-kB-p65 in vivo and in vitro
↓NF-Κb-p65 in vivo and in vitro [69]
[69]
↓NF-Κb-p65 in vivo and
↓NF-Κb-p65
colon
↓NF-Κb-p65
body
colon
MLCK,
MLCK, ↑MDA, ↓GSH, ↓SOD, ↑iNOS,
SOD, ↑iNOS, ↑COX-2 in vivo
SOD, ↑iNOS, ↑
treated cells
in vitro
weight
in vitro
weight
↑p-p65 nuclear translocation in NF-κB p65
↓ ↑hPXR via binding to hPXR-
↑p-p65 nuclear translocation in vitro and ↑NF-kB p65 phosphorylation, ↑IκBα phosphorylation/degradation, ↑iNOS expression, ↑ICAM-1 expression, ↑MCP-1 expression, ↑COX-2 expression, ↑TNF-α expression, ↑IFN-γ expression, ↑IL-6 expression, ↑MPO, ↑ NO, ↑PGE2, ↑TNF-α, ↑IL-6 in vivo
↓p-p65 nuclear translocation, hPXR via binding to hPXRLBD in vitro and ↓NF-κB p65 phosphorylation, ↓IκBα phosphorylation/degradation, ↓iNOS expression, ↓ICAM-1 expression, COX-2 expression, ↓TNF-α expression, ↓IFN-γ expression and ↓IL-6 expression in vivo
p-p65 nuclear translocation in NF-κB p65
↓p-p65 nuclear translocation, ↑hPXR via binding to hPXR-LBD in vitro and ↓NF-kB p65 phosphorylation, ↓IκBα phosphorylation/degradation, ↓iNOS expression, ↓ICAM-1 expression, ↓COX-2 expression, ↓TNF-α expression, ↓IFN-γ expression and ↓IL-6 expression in vivo
↓p-p65 nuclear translocation, ↑hPXR via binding to hPXRLBD ↓NF-κB p65 phosphorylation, ↓IκBα phosphorylation/degradation, ↓iNOS expression, ↓ICAM-1 expression, COX-2 expression, ↓TNF-α expression, ↓IFN-γ expression and ↓IL-6 expression in vivo
p-p65 nuclear translocation in NF-κB p65
↓p-p65 nuclear translocation, ↑hPXR via binding to hPXR-
p-p65 nuclear translocation in NF-κB p65
↑IκBα phosphorylation/degradation, ↑iNOS expression, ↑ICAM-1 expression, MCP-1 expression, ↑COX-2 expression, ↑TNF-α expression, IFN-γ expression,
and ↓NF-κB p65 phosphorylation, ↓IκBα phosphorylation/degradation, ↓iNOS expression, ↓ICAM-1 expression, COX-2 expression, ↓TNF-α expression, ↓IFN-γ expression and ↓IL-6 expression in vivo
↑IκBα phosphorylation/degradation, ↑iNOS expression, ↑ICAM-1 expression, MCP-1 expression, ↑COX-2 expression, ↑TNF-α
IκBα phosphorylation/degradation, ↑iNOS expression, ↑ICAM-1 expression, MCP-1 expression, ↑COX-2 expression, ↑TNF-α
↑IκBα phosphorylation/degradation, ↑iNOS expression, ↑ICAM-1 expression, MCP-1 expression,
and ↓NF-κB p65 phosphorylation, ↓IκBα phosphorylation/degradation, ↓iNOS expression, ↓ICAM-1 expression, COX-2 expression, ↓TNF-α expression, ↓IFN-γ expression and ↓IL-6 expression in vivo
Alantolactone C57BL/6 mice model of colitis and LPS-induced RAW 264.7 and HT29 colorectal treated
↓bloody diarrhea, colon length, ↑histological injury, ↑inflammatory cell
Alantolactone C57BL/6 mice model of colitis and LPS-induced RAW 264.7 and HT29 colorectal treated
↓bloody diarrhea, colon length, ↑histological injury, ↑inflammatory cell
Alantolactone
DSS-induced C57BL/6 mice model of colitis and LPS-induced RAW 264.7 and HT-29 colorectal treated cells
↓Body weight, ↓bloody diarrhea, ↓colon length, ↑histological injury, ↑inflammatory cell infiltration
mice model of colitis and LPS-induced RAW 264.7 and HT29 colorectal treated
colon length, ↑histological injury, ↑inflammatory cell
and LPS-induced RAW 264.7 and HT29 colorectal treated cells
[70]
histological injury, ↑inflammatory cell infiltration
incubated for 2 h in vitro 50 mg/kg/day
in vitro 50 mg/kg/day orally/9 days in
0–25 µM incubated for 2 h in vitro 50 mg/kg/day orally/9 days in vivo
mg/kg/day orally/9 days in vivo
in vitro 50 mg/kg/day orally/9 days in
[70]
[70]
[70]
[70]
COX-2 expression, ↑TNF-α
IFN-γ expression, ↑MPO, ↑ NO, IL-6 in vivo
↑MPO, ↑ NO, IL-6 in vivo
IFN-γ expression,
IFN-γ expression, ↑MPO, ↑ NO, IL-6 in vivo
MPO, ↑ NO, IL-6 in vivo
NF-κBp65, ↑TLR4,
↓MyD88 expression, ↓NFκBp65 expression, ↓TLR expression, ↑SIGIRR
NF-κBp65, ↑TLR4, SIGIRR expression, ↑TLR/NF-
↓MyD88 expression, ↓NF-
SIGIRR expression, ↑TLR/NFκB
NF-κBp65, ↑TLR4, ↓SIGIRR expression, ↑TLR/NF-
↓MyD88 expression, ↓NFκBp65 expression, ↓TLR [71]
NF-κBp65, ↑TLR4, ↓SIGIRR expression, ↑TLR/NF-
MyD88 expression, κBp65 expression, ↓TLR [71]
↑DAI, ↓colon length,
DSS-induced BALB/c30, 90, 270, 180,
Sinomenine intake, ↑pasty stools,
mice model of colitis 540 mg/kg/day
intake, ↑pasty stools,
Sinomenine
15 of 44
15 of 44
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
Mechanisms in Regulation of NF-kB in IBD
Ref.
Phytochemicals
Sinomenine
TLR/NF-κB,
↓MyD88 expression, ↓NF-kBp65 expression, ↓TLR expression, ↑SIGIRR expression, ↓TLR/NF-kB, ↓expression of IFN-γ, IL-1β, TNF-α, IL-6, and IL-12
TLR/NF-κB, ↓expression of IFN-γ, IL-1β, TNF-α, IL-6, and IL-12
inflammatory cell κB, ↓expression of IFN-γ, IL-1β, TNF-α
expression of IFN-γ, IL-1β, TNF-α, IL-6, and IL-12
↓Body weight, ↓food intake, ↑pasty stools, ↑DAI, ↓colon length, and ↑inflammatory cell infiltration
orally/9 days
30, 90, 270, 180, 540 mg/kg/day and 1.6 g/kg/day orally/9 days
↑MyD88, ↑NF-kBp65, ↑TLR4, ↓SIGIRR expression, ↑TLR/NF-kB
DSS-induced BALB/c mice model of colitis
[71]
↓NF-κB p65, ↑PPARγ, ↑PPARγ NF-κB mRNA, ↓IL-1β IL-6 mRNA, ↓TNF-α
↓NF-κB p65, ↑PPARγ, ↑PPARγ
↓ ↑PPARγ, ↑PPARγ
↓NF-kB p65, ↑PPARγ, ↑PPARγ mRNA, ↓NF-kB mRNA, ↓IL-1β mRNA, ↓IL-6 mRNA, ↓TNF-α mRNA, ↓p-IκBα, ↑PPARγ siRNA in vitro and ↓nuclear NF-kB-p65, ↑PPARγ expression, ↓nuclear translocation of NF-kB-p65, ↓cytoplasmic p-IκBα expression, ↑PPARγ mRNA and ↓NF-kB mRNA in vivo
NF-κB mRNA, ↓IL-1β mRNA, ↓IL-6 mRNA, ↓TNF-α mRNA, ↓p-IκBα, ↑PPARγ siRNA in vitro and ↓nuclear NF-κB-p65, PPARγ expression, nuclear translocation of NF-κB-p65, ↓cytoplasmic p-IκBα
iNOS, ↑ IL1β, ↑ IL-6, ↑TNF-α, ↑NF-κB-p65 and ↓PPARγ in vitro and ↑COX-2, ↑iNOS, IL-1β, ↑IL-6, ↑TNF-α, ↑nuclear NF-κB-p65, ↓PPARγ protein in vivo
mRNA, ↓ ↓TNF-α mRNA, κBα, ↑PPARγ
Convallatoxin
mRNA, ↓p-IκBα, ↑PPARγ siRNA in vitro and ↓nuclear NF-κB-p65, PPARγ expression, nuclear translocation of NF-κB-p65, ↓cytoplasmic p-IκBα
↑NF-kB, ↑COX-2, ↑iNOS, ↑ IL-1β, ↑ IL-6, ↑TNF-α, ↑NF-kB-p65 and ↓PPARγ in vitro and ↑COX-2, ↑iNOS, ↑IL-1β, ↑IL-6, ↑TNF-α, ↑nuclear NF-kB-p65, ↓PPARγ protein in vivo
↑NF-κB, ↑COX-2, ↑iNOS, ↑ IL1β, ↑ IL-6, ↑TNF-α, ↑NF-κB-p65 and ↓PPARγ in vitro and
Convallatoxin
↑NF-κB, ↑COX-2, ↑iNOS, ↑ β, ↑ TNF-α, ↑NF-κ and ↓PPARγ in vitro and ↑COX-2, ↑iNOS, IL-1β, ↑IL-6, ↑TNF-α, ↑nuclear NF-κB-p65, ↓PPARγ in vivo
↓Colon length, ↑colon and spleen weights, ↓body weight, ↑inflammatory cell infiltration, ↑ulceration, ↑necrosis, ↑congestion and ↑edema, ↑IL-1β, IL-6, and TNF-α in the colon
body weight, ↑inflammatory cell infiltration, ulceration, ↑necrosis, congestion and ↑ ↑IL-1β, IL6, and TNF-α in the
body weight, ↑inflammatory cell infiltration, ↑
inflammatory cell infiltration, ulceration, ↑necrosis, congestion and ↑ ↑IL-1β, IL6, and TNF-α in the
LPS-induced RAW264.7 and BMDMs macrophages and DSS-induced C57BL/6 mice model of colitis
10–50 nM incubated for 12 or 24 h in vitro and 50 or 150 µg/kg/day orally/10 days in vivo
BMDMs macrophages and DSS-induced C57BL/6 mice model
24 h in vitro and 50 or 150 µg/kg/day orally/10 days in
in vitro and ↓nuclear
in vitro and
macrophages and DSS-induced C57BL/6 mice model
50 or 150 µg/kg/day orally/10 days in
[72]
[72]
NF-κB-p65, PPARγ expression, nuclear
[72]
↑COX-2, ↑iNOS, IL-1β, ↑IL-6, ↑TNF-α, ↑nuclear NF-κB-p65, ↓PPARγ protein in vivo
DSS-induced
↑ ↑IL-1β
κB-p65, κBα
in
PPARγ mRNA NF-κB mRNA in vivo
PPARγ mRNA NF-κB mRNA in vivo
PPARγ
↑Nitrites, ↑TNF-α, ↑IL-1β, ↑IL-
↑Nitrites, ↑TNF-α, ↑IL-1β, ↑ ↑
↑NF-kB-p65 nuclear translocation, ↑IkBα phosphorylation and
↑Nitrites, ↑TNF-α, ↑IL-1β, ↑IL-6, ↑COX-2, ↑iNOS, ↑NF-kB-p65 nuclear translocation, ↑IkBα phosphorylation and degradation in vitro and ↑MPO, ↑TNF-α, ↑IL-1β, ↑IL-6, ↑Nitrites, ↑COX-2, ↑iNOS, ↑nuclear NF-kB (p65), ↑phosphorylation of IκBα (p-IκBα/IκBα), ↑NF-kB (p65)-DNA binding activity, ↑p-p38/p38, ↑p-ERK/ERK, ↑Akt phosphorylation, ↓GSH, and ↑TBARS in vivo
↑NF-kB-p65 nuclear translocation, ↑IkBα phosphorylation and
↑IkBα phosphorylation and
↓NF-Kb-p65e expression, ↓IkBα phosphorylation and degradation in vitro and ↓nuclear NF-κB (p65), ↓phosphorylation of IκBα (pIκBα/IκBα), NF-κB (p65)-DNA binding activity,↓p-p38/p38,
↓NF-kB-p65e expression, ↓IkBα phosphorylation and degradation in vitro and ↓nuclear NF-kB (p65), ↓phosphorylation of IκBα (p-IκBα/IκBα), ↑NF-kB (p65)-DNA binding activity, ↓p-p38/p38, ↑p-ERK/ERK, and ↓Akt phosphorylation in vivo
↓NF-Kb-p65e expression, ↓IkBα phosphorylation and degradation in vitro and ↓nuclear NF-κB (p65), ↓phosphorylation of IκBα (pIκBα/IκBα), NF-κB (p65)-DNA binding activity,↓p-p38/p38,
↓ ↓IkBα phosphorylation and degradation in vitro and ↓nuclear NF-κB (p65), ↓phosphorylation of IκBα (pIκBα/IκBα NF-κ b ↓p-p38/p38, ↑ ↓Akt
in vitro and ↑MPO,
in vitro and ↑MPO, ↑TNF-α, IL-1β, IL-6, ↑Nitrites, ↑COX-2, ↑iNOS, ↑nuclear NF-κB (p65),
in vitro and ↑MPO, ↑TNF-α, IL-1β, IL-6, ↑Nitrites, ↑COX-2, ↑iNOS, ↑nuclear NF-κB (p65),
Fisetin
IL-6, ↑Nitrites, ↑COX-2, ↑iNOS, ↑nuclear NF-κB (p65), ↑phosphorylation of IκBα (p-
Fisetin
↑DAI, body weight, ↓colon length, ↓crypts, ↓goblet cells,
Fisetin DSS-induced Balb/C mice model of colitis and LPS-induced macrophages treated
mg/kg/day orally/8 days in
↑ body weight, ↓ ↓ ↓
mg/kg/day
↓colon length, ↓crypts, ↓goblet cells, ↑inflammatory cell
DSS-induced Balb/C mice model of colitis and LPS-induced macrophages treated cells
5 and 10 mg/kg/day orally/8 days in vivo and 0–50 µM incubated for 24 h in vitro
↑DAI, ↓body weight, ↓colon length, ↓crypts, ↓goblet cells, ↑inflammatory cell infiltration,
mice model of colitis and LPS-induced macrophages treated cells
orally/8 days in
mice model of colitis
in vivo and
[73]
vivo and 0–50 µM incubated for 24 h
[73]
[73]
vivo and 0–50 µM
inflammatory cell
phosphorylation of IκBα (pNF-κB (p65)-
inflammatory cell
phosphorylation of IκBα (pNF-κ
NF-κB (p65)-
↑p-ERK/ERK, and ↓Akt phosphorylation in vivo
↑p-ERK/ERK, and ↓Akt phosphorylation in vivo
p-p38/p38,
p-p38/p38, Akt
p-p38/p38, Akt
in vivo
GSH, and ↑TBARS in vivo
↓GSH, and
↓
Body weight, colon Oxide-nitrosative stress, Oxide-nitrosative stress,
Metabolites 2023, 13, 96 19 of 47
↓phosphorylation of IκBα (p-
and LPS-induced vivo and ↓goblet cells, ↑nuclear NF-κB (p65),
[73]
Table 1. Cont.
binding activity, ↑p-p38/p38, p-ERK/ERK, phosphorylation, GSH, and
phosphorylation in vivo
Related Molecular Mechanisms in Regulation of NF-kB in IBD
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
Ref.
Phytochemicals
Genipin
Genipin
TNF-α and ↓IL-1β expression, ↓NF-kB signaling, ↑Nrf2 signaling and ↑HO-1 expression
↓Body weight, ↑intestinal
↓MPO, ↓MDA, ↓TNF-α and ↓IL-1β expression, ↓NF-kB signaling, ↑Nrf2 signaling and ↑HO-1 expression
↑MPO, ↑MDA, TNF-α, IL-1β, ↑NF-kB signaling, ↓Nrf2 signaling and HO-1
DSS-induced C57BL/6 mice model of colitis
2.5, 5, 10 mg/kg/day orally/14 days
intestinal epithelial destruction, ↑crypt abscesses, and ↑goblet
↑MPO, ↑MDA, ↑TNF-α, ↑IL-1β, ↑NF-kB signaling, ↓Nrf2 signaling and ↓HO-1
DSS-induced C57BL/6 mice model of colitis
2.5, 5, 10 mg/kg/day orally/14 days
epithelial destruction, ↑crypt abscesses, and ↑goblet cells loss
[74]
[74]
, 96
Metabolites 2023, 13, 96 16 of 44
, 13
Metabolites 2023, 13, 96 16 of 44
↓Oxide-nitrosative stress, ↓iNOS, ↓TNF-α, ↓IL-1β, ↓IFN-γ, ↓COX-2 mRNA, ↓LTB4 and ↓IkBα expression levels, ↓NF-kB signaling, ↑occludin, ↑claudin-1, ↑zonula occludens-1 and ↑IL-10 expression levels and ↓caspase-1
Oxide-nitrosative stress,
↑Oxide-nitrosative stress, ↑iNOS, ↑TNF-α, ↑IL-1β, ↑IFN-γ, ↑COX-2 mRNA, ↑LTB4, ↑IkBα, ↑NF-kB signaling, ↓occludin, ↓claudin-1, ↓zonula occludens-1, ↑caspase-1 and ↓IL-10
↓iNOS, ↓TNF-α, ↓IL-1β, ↓IFNγ, ↓COX-2 mRNA, ↓LTB4 and
IL-1β, ↑IFNCOX-2 mRNA, ↑LTB4,
[75]
Piperine
↑IkBα, ↑NF-kB signaling, ↓occludin, ↓claudin-1, ↓zonula occludens-1, caspase-1 and
↓IkBα expression levels, ↓NFkB signaling, occludin, ↑claudin-1, zonula occludens1 and ↑IL-10 expression levels and ↓
↑IkBα, ↑ ↓occludin, ↓claudin-1, ↓
↓IkBα ↓NF-
↓Body weight, ↑colon weight-to-length ratio, and ↑ulceration
TNBS-induced Sprague–Dawley mice model of colitis
↑IkBα, ↑ ↓occludin, ↓claudin-1, ↓
↓IkBα ↓NF-
↑IkBα, ↑NF-kB signaling, ↓occludin, ↓claudin-1, ↓zonula
↓IkBα expression levels, ↓NFkB signaling, occludin, ↑claudin-1, zonula occludens1 and ↑IL-10 expression levels
10, 20, and 40 mg/kg/day orally/14 days
[75]
occludin, ↑claudin-1,
occludin, ↑claudin-1,
caspase-1 and
↑
↑
↓MPO, ↓iNOS, ↓TNF-α, ↓IL-1β, IL-6, ↓ MIP-1α and ↓IL-
Ligustilide
↓
↓Body weight and
↓MPO, ↓iNOS, ↓TNF-α, ↓IL-1β, IL-6, ↓IL-12, MIP-1α and ↓IL-
↓
↓ ↓ ↓TNF-α, ↓IL-1β, ↓IL-6, ↓IL-12, MIP-1α and ↓IL-
↓TNF-α, ↓IL-1β, ↓IL-6, ↓IL-12, MIP-1α and ↓IL-
Ligustilide
Ligustilide
↓MPO, ↓iNOS, ↓TNF-α, ↓IL-1β, ↓IL-6, ↓IL-12, ↓MIP-1α and ↓IL-17 expression levels, ↑PPARγ expression and signaling and ↓NF-kB-p65 expression
↑MPO, ↑iNOS, TNF-α, IL-1β, ↑IL-6, ↑IL-12, MIP-1α, ↑IL-17, ↓PPARγ and ↑NF-kB-p65
↓Body weight and ↓colon length, ↑diarrhea, ↑rectal bleeding, ↑ulceration, and ↑inflammatory cells infiltration
colon length, ↑diarrhea, rectal bleeding, ulceration,
↑MPO, ↑iNOS, TNF-α, ↑IL-1β, ↑IL-6, ↑IL-12, MIP-1α, ↑ ↓PPARγ and ↑
↑MPO, ↑iNOS, ↑TNF-α, ↑IL-1β, ↑IL-6, ↑IL-12, ↑MIP-1α, ↑IL-17, ↓PPARγ and ↑NF-kB-p65
↑MPO, ↑iNOS, TNF-α, ↑IL-1β, ↑IL-6, ↑IL-12, MIP-1α, ↑ ↓PPARγ and
↑MPO, ↑iNOS, TNF-α, ↑IL-1β, ↑IL-6, ↑IL-12, MIP-1α, ↑IL-17, ↓PPARγ and NF-kB-p65
rectal bleeding, ulceration, inflammatory cells
17 expression levels, ↑PPARγ expression and signaling and ↓NF-kB-p65 expression
↑diarrhea, bleeding,
C57BL/6 mice model of colitis
[76]
mg/kg/day orally/14 days
↑PPARγ
C57BL/6 mice model of colitis
mg/kg/day
DSS-induced C57BL/6 mice model of colitis
15, 30, and 60 mg/kg/day orally/14 days
↑PPARγ
17 expression levels, ↑PPARγ expression and signaling and ↓NF-kB-p65 expression
C57BL/6 mice model of colitis
mg/kg/day
C57BL/6 mice model of colitis
[76]
mg/kg/day orally/14 days
[76]
bleeding,
inflammatory cells
↓
↓
Evodiamine
IL-1β, ↑IL-6, ↑p-NF-kB p65, p-IkB, ↑NLRP3, ↑ASC, ↑caspase-1, ↓ZO-1 and ↓occludin
↓MPO, ↓ ↓IL-1β, ↓IL-6, ↓p-NF-kB p65, ↓p-IkB, ↓NLRP3, ↓ASC, ↓caspase-1, ↑ZO-1 and ↑occludin
Evodiamine
Evodiamine
IL-1β, ↑IL-6, ↑p-NF-kB p65, p-IkB, ↑NLRP3, ↑ASC, ↑caspase-1, ↓ZO-1 and ↓occludin
↓MPO, ↓TNF-α, ↓IL-1β, ↓IL-6, ↓p-NF-kB p65, ↓p-IkB, ↓NLRP3, ↓ASC, ↓caspase-1, ↑ZO-1 and ↑occludin
↑MPO, ↑TNF-α, ↑IL-1β, ↑IL-6, ↑p-NF-kB p65, ↑p-IkB, ↑NLRP3, ↑ASC, ↑caspase-1, ↓ZO-1 and ↓occludin
↓MPO, ↓TNF-α, ↓IL-1β, ↓IL-6, ↓p-NF-kB p65, ↓p-IkB, ↓NLRP3, ↓ASC, ↓caspase-1, ↑ZO-1 and ↑occludin
↑MPO, TNF-α, IL-1β, ↑IL-6, ↑p-NF-kB p65, p-IkB, ↑NLRP3, ↑ASC, ↑ ↓ZO-1 and ↓
↓MPO, ↓TNF-α, ↓IL-1β, ↓IL-6, ↓p-NF-kB p65, ↓p-IkB, ↓NLRP3, ↓ASC, ↓caspase-1, ↑ZO-1 and
↑MPO, TNF-α, IL-1β, ↑IL-6, ↑p-NF-kB p65, p-IkB, ↑NLRP3, ↑ASC, ↑ ↓ZO-1 and ↓
↓MPO, ↓TNF-α, ↓IL-1β, ↓IL-6, ↓p-NF-kB p65, ↓p-IkB, ↓NLRP3, ↓ASC, ↓caspase-1, ↑ZO-1 and
fecal bleeding, colon
fecal bleeding, colon shortening, and ↓body
↑Diarrhea, ↑fecal bleeding, ↑colon shortening, and ↓body weight
bleeding, colon shortening, and ↓body
DSS-induced C57BL/6 mice model mg/kg/day
DSS-induced C57BL/6 mice model
20, 40, and 80 mg/kg/day orally/10 days
bleeding, colon
DSS-induced C57BL/6 mice model of colitis
20, 40, and 80 mg/kg/day orally/10 days
C57BL/6 mice model of colitis
mg/kg/day orally/10 days
[77]
C57BL/6 mice model of colitis
mg/kg/day orally/10 days
[77]
[77]
↓
↓
orally/10 days
↑p-65, ↑IkBα phosphorylation
↓p-65, ↓IkBα phosphorylation and degradation, ↓NF-kB nuclear translocation, ↓iNOS mRNA, ICAM-1 mRNA, ↓MCP-1 mRNA, ↓COX-2 mRNA, ↓TNF-α mRNA, ↓IL-6 mRNA, and MPO
↑ ↑IkBα phosphorylation
↓ ↓IkBα phosphorylation and degradation, ↓NF-kB nuclear translocation, ↓iNOS mRNA, ↓ICAM-1 mRNA, ↓MCP-1 mRNA, ↓COX-2 mRNA, ↓TNF-α mRNA, ↓IL-6 mRNA, and MPO
↑p-65, ↑IkBα
↓p-65, ↓IkBα
IkBα phosphorylation
IkBα phosphorylation and degradation, ↓NF-kB nuclear translocation, ↓iNOS mRNA, ↓ICAM-1 mRNA, ↓MCP-1 mRNA, ↓COX-2 mRNA, ↓TNF-α mRNA, ↓IL-6 mRNA, and MPO ↓IL-1β mRNA, ↓TNF-α mRNA,
IkBα phosphorylation
IkBα phosphorylation and degradation, ↓NF-kB nuclear translocation, ↓iNOS mRNA, ↓ICAM-1 mRNA, ↓MCP-1 mRNA, ↓COX-2 mRNA, ↓TNF-α mRNA, ↓IL-6 mRNA, and MPO
↑NF-kB nuclear translocation, ↑iNOS mRNA, ↑ICAM-1 mRNA, ↑MCP-1 mRNA, COX-2 mRNA, ↑TNF-α mRNA, ↑IL-6
↑NF-kB nuclear translocation, ↑iNOS mRNA, ↑ICAM-1 mRNA, ↑MCP-1 mRNA, COX-2 mRNA, ↑TNF-α mRNA, ↑IL-6
phosphorylation and degradation, ↑NF-kB nuclear translocation,
phosphorylation and degradation, ↓NF-kB nuclear translocation,
↑NF-kB nuclear translocation, ↑iNOS mRNA, ↑ICAM-1 mRNA, ↑MCP-1 mRNA, COX-2 mRNA, ↑TNF-α mRNA, ↑IL-6
↑NF-kB nuclear translocation, ↑iNOS mRNA, ↑ICAM-1 mRNA, ↑MCP-1 mRNA, COX-2 mRNA, ↑TNF-α mRNA, ↑IL-6
Chrysin
Chrysin
fecal bleeding, crypt distortion, and
fecal bleeding,
↓Body weight, ↑diarrhea, ↑fecal bleeding, ↑crypt distortion, and ↑inflammatory exudate
fecal bleeding,
fecal bleeding, crypt distortion, and
C57BL/6 mice model of colitis
[78]
25 mg/kg/day orally/10 days
TNBS-induced C57BL/6 mice model of colitis
25 mg/kg/day orally/ 10 days
C57BL/6 mice model of colitis
orally/10 days
C57BL/6 mice model of colitis
C57BL/6 mice model of colitis
[78]
↑iNOS mRNA, ↑ICAM-1 mRNA, ↑MCP-1 mRNA, ↑COX-2 mRNA, ↑TNF-α mRNA, ↑IL-6 mRNA, and ↑MPO
↓iNOS mRNA, ↓ICAM-1 mRNA, ↓MCP-1 mRNA, ↓COX-2 mRNA, ↓TNF-α mRNA, ↓IL-6 mRNA, and ↓MPO
[78]
orally/10 days
orally/10 days
inflammatory exudate
MPO
MPO
MPO
MPO
↓TNF-α mRNA, ↓ ↓
IL-1β mRNA, ↓TNF-α mRNA,
↓IL-1β mRNA, ↓TNF-α mRNA,
↑
- Metabolites 2023, 13, 96 20 of 47
of colitis orally/10 days
weight ↓occludin ↑occludin
Table 1. Cont.
TNBS-induced
25 mg/kg/day orally/10 days
nuclear translocation, ↑iNOS mRNA, ↑ICAM-1 mRNA, ↑MCP-1 mRNA, COX-2
↑diarrhea, ↑fecal bleeding, distortion, and
nuclear translocation, ↓iNOS mRNA, ICAM-1 mRNA, ↓MCP-1 mRNA, COX-2
TNBS-induced C57BL/6 mice model of colitis
↑ distortion, and ↑inflammatory exudate
↑ ↑MCP-1 mRNA, mRNA, ↑TNF-α mRNA, ↑IL-6 mRNA, and MPO
↓ICAM-1 mRNA, ↓MCP-1 mRNA, COX-2 mRNA, ↓TNF-α mRNA, ↓IL-6 mRNA, and MPO
[78]
Related Molecular Mechanisms in Regulation of NF-kB in IBD
[78]
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
of colitis
orally/10 days
Ref.
Phytochemicals
↑ mRNA, and ↑MPO
↓IL-6 mRNA, and ↓MPO
↓IL-1β mRNA, ↓TNF-α mRNA, ↓IL-6 mRNA, ↓NF-kB nuclear translocation, ↓IkBa phosphorylation, ↓phosphorylation of p65, ↓NF-kB DNA binding activity, ↓NLRP3 mRNA and ↓pro-caspase-1 mRNA in vitro and ↓NF-kB, ↓NF-kB-p65, ↓IkBa phosphorylation, ↓p65, ↓p65 phosphorylation and ↓NF-kB DNA binding activity in vivo
↓TNF-α mRNA, NF-kB nuclear IkBa
↓TNF-α mRNA, NF-kB nuclear
translocation, IkBa phosphorylation, ↓phosphorylation of p65, ↓NFkB DNA binding activity, ↓NLRP3 mRNA and ↓procaspase-1 mRNA in vitro and ↓NF-kB, NF-kB-p65, ↓IkBa phosphorylation, ↓p65, ↓p65 phosphorylation and ↓NF-kB DNA binding activity in vivo
↑NLRP3 mRNA and ↑procaspase-1 mRNA in vitro and ↑IL-1β, IL-6, ↑NF-kB p65, ↑cleaved caspase-1 (p10), ↑cleaved-IL-1β, NLRP3 and ↑ASC in vivo
phosphorylation, ↓phosphorylation of p65, ↓NFkB DNA binding activity, ↓NLRP3 mRNA and ↓procaspase-1 mRNA in vitro and ↓NF-kB, ↓NF-kB-p65, ↓IkBa phosphorylation, ↓p65, ↓p65 phosphorylation and ↓NF-kB DNA binding activity in vivo
↓Body weight, ↓colon length, ↓spleen weight, ↑inflammatory cell infiltration, ↑ulcers, ↑edema and ↑congestion, ↑CD11b+ F4/80+ macrophages and ↑CD11b+ Gr-1+ neutrophils,
↑NLRP3 mRNA and ↑pro-caspase-1 mRNA in vitro and ↑IL-1β, ↑TNF-α, ↑IL-6, ↑NF-kB p65, ↑cleaved caspase-1 (p10), ↑cleaved-IL-1β, ↑NLRP3 and ↑ASC in vivo
spleen weight, ↑inflammatory cell infiltration, ulcers, ↑edema and
inflammatory cell infiltration, ulcers, ↑edema and ↑congestion, ↑CD11b+ F4/80+ macrophages
↑NLRP3 mRNA and ↑procaspase-1 mRNA in vitro and ↑IL-1β, ↑TNF-α, IL-6, ↑NF-kB p65, ↑cleaved caspase-1 (p10), ↑cleaved-IL-1β, NLRP3 and ↑ASC in vivo
Wogonoside
Wogonoside mice model of colitis and LPS-induced Human acute monocytic leukemia
DSS-induced BALB/c mice model of colitis and LPS-induced Human acute monocytic leukemia
DSS-induced BALB/c mice model of colitis and LPS-induced Human acute monocytic leukemia THP-1 treated cells
mg/kg/day orally/10 days in
Wogonoside
12.5, 25, or 50 mg/kg/day orally/10 days in vivo and 0.1 mM incubated for 4 h in vitro
mg/kg/day orally/10 days in vivo and 0.1 mM
[79]
[79]
and 0.1 mM incubated for 4 h
[79]
↑CD11b+ F4/80+ macrophages
CD11b+ Gr-1+
↓Body weight, ↓colon
Oxymatrine
↓ZO-1 mRNA, ↓ mRNA, ↓claudin-2 mRNA, ↑IL6, ↑TLR9, ↑Myd88 and ↑p-NFκB P65
↓ ↓TNF-α mRNA, ↓IL-6 mRNA, NF-kB, ↓TLR9 expression, ↓Myd88, ↓TLR9/Myd88/NF-κB pathway
↓colon
↓IL-1β mRNA, ↓TNF-α mRNA, ↓IL-6 mRNA, ↓NF-kB, ↓TLR9 expression, ↓Myd88, ↓TLR9/Myd88/NF-kB pathway
Oxymatrine
Oxymatrine
↑ulcers, ↓goblet cells, and ↑inflammatory cell
↓Body weight, ↓colon length, ↑DAI, ↑ulcers, ↓goblet cells, and ↑inflammatory cell infiltration
↓ZO-1 mRNA, occludin mRNA, ↓claudin-2 mRNA, ↑IL6, ↑TLR9, Myd88 and ↑p-NFκB P65
IL-1β mRNA, ↓TNF-α mRNA, ↓IL-6 mRNA, NF-kB, ↓TLR9 expression, Myd88,
↓ZO-1 mRNA, ↓occludin
17 of 44
↑ulcers, ↓goblet cells, and ↑inflammatory cell
[80]
TNBS-induced rats model of colitis
mg/kg/day intraperitoneally/7
mRNA, ↓claudin-2 mRNA, ↑IL-6, ↑TLR9, ↑Myd88 and ↑p-NF-kB P65
TNBS-induced rats model [80] of colitis
10, 30, or 60 mg/kg/day intraperitoneally/7 days
model of colitis peritoneally/7 days
[80]
TLR9/Myd88/NF-κB pathway
DSS-induced C57BL/6J mice model of colitis and LPS-induced RAW264.7 and IEC-6
Epicatechin
Epicatechin
Body weight, ↓colon length, ↑intestinal bleeding, DAI, and ↑CMDI scores
DSS-induced C57BL/6J mice model of colitis and LPS-induced RAW264.7 and IEC-6 treated cells
100, 200, or 300 mg/kg/day orally/7 days in vivo and 0.1 µM, 1 µM or 10 µM incubated for 4 h in vitro
↓Body weight, ↓colon length, ↑intestinal bleeding, ↑DAI, and ↑CMDI scores
orally/7 days in vivo and 0.1 µM, 1 µM or 10 µM incubated for 4 h in vitro
↑TNF-α, ↑ NO, ↑MPO and ↑
↑TNF-α, ↑IL-6, ↑NO, ↑MPO and ↑NF-kB ↓NF-kB expression [81]
↓NF-kB expression
↑ ↑
- Metabolites 2023, 13, 96 21 of 47
Epicatechin
↓Body weight, ↓colon
C57BL/6J mice
100, 200, or 300
orally/7 days in
C57BL/6J mice
model of colitis and LPS-induced
Table 1. Cont.
RAW264.7 and IEC-6 treated cells
bleeding, ↑DAI, and and ↑NF-kB
↓NF-kB expression [81]
vivo and 0.1 µM, 1 µM or 10 µM incubated for 4 h
µM or 10 µM incubated for 4 h
Related Molecular Mechanisms in Regulation of NF-kB in IBD
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
↑CXCL-1 mRNA, IL-8 mRNA and COX-2 mRNA in vitro and ↑IL-1β expression, ↑TNF-α
↑CMDI scores
RAW264.7 and IEC-6 treated cells
treated cells
Ref.
Phytochemicals
CXCL-1 mRNA, IL-8 mRNA, and COX-2 mRNA, ↑PPAR-γ expression both at protein and
↑DAI, ↑inflammatory
↑CXCL-1 mRNA, ↑IL-8 mRNA
mg/kg/day orally
Thymoquinone
and COX-2 mRNA in vitro and ↑TNF-α IL-6, expression
↑CXCL-1 mRNA, ↑IL-8 mRNA and COX-2 mRNA in vitro and ↑IL-1β expression, ↑TNF-α expression, ↑IL-6, expression ↑IL-6 mRNA, ↑IL-1β mRNA, ↑TNF-α mRNA, ↑COX-2, ↑iNOS, ↑COX-2 mRNA, ↑iNOS mRNA, ↑p-ERK, ↑p-JNK, ↑p-p38, ↑phosphorylation of the NF-kB protein and ↓PPAR-γ expression in vivo
IL-6, expression IL-1β mRNA,
↓CXCL-1 mRNA, IL-8 mRNA, and COX-2 mRNA, ↑PPAR-γ expression both at protein and mRNA and ↓IL-6 mRNA, ↓IL-1β mRNA, ↓TNF-α mRNA, ↓p-ERK, ↓p-JNK, ↓pp38, ↓phospho-NF-κB protein and ↑PPAR-γ in vivo
in vitro and ↑TNF-α
↓CXCL-1 mRNA, IL-8 mRNA, and COX-2 mRNA, ↑PPAR-γ expression both at protein and mRNA in vitro and ↓IL-6 mRNA, ↓IL-1β mRNA, ↓TNF-α mRNA, ↓p-ERK, ↓p-JNK, ↓pp38, ↓phospho-NF-κB protein and ↑PPAR-γ in vivo
↓CXCL-1 mRNA, IL-8 mRNA, and COX-2 mRNA, ↑PPAR-γ expression both at protein and mRNA in vitro and ↓IL-6 mRNA, ↓IL-1β mRNA, ↓TNF-α mRNA,
and ↓IL-6 mRNA, ↓IL-1β mRNA, ↓TNF-α mRNA, ↓ ↓p-JNK, ↓pp38, ↓phospho-NF-κB protein and ↑PPAR-γ in vivo
[82]
COX-2, ↑iNOS, ↑COX-2 mRNA, ↑iNOS
mg/kg/day orally fo/r 8 days in vivo
DAI,
Thymoquinone
fo/r 8 days in vivo and 0, 12.5, 50, 100, 150, and 200 µM incubated for 24 h in vitro
cells infiltration, ↓crypts, villi,
↑submucosal edema, ↑epithelium
↑IL-6, expression ↑IL-6 mRNA, ↑IL-1β mRNA, ↑TNF-α mRNA, COX-2, ↑iNOS, COX-2 mRNA, ↑iNOS mRNA, p-JNK, ↑p-
0, 12.5, 50, 100, 150, and 200 µM incubated for 24 h in vitro
Thymoquinone
20 and 40 mg/kg/day orally for 8 days in vivo and 0, 12.5, 50, 100, 150, and 200 µM incubated for 24 h in vitro
IL-6 mRNA, IL-1β mRNA, ↑TNF-α mRNA, COX-2, ↑iNOS, ↑COX-2 mRNA, ↑iNOS mRNA, p-JNK, ↑p-
cells infiltration, ↑MPO, villi, ↑submucosal edema, and ↑epithelium
TNF-α-induced HT29 treated cells
↑DAI, ↑inflammatory cells infiltration, ↑MPO, ↓crypts, ↓villi, ↑submucosal edema, and ↑epithelium destruction
DSS-induced C57BL/6J mice model of colitis and TNF-α-induced HT-29 treated cells
C57BL/6J mice model of colitis and TNF-α-induced HT29 treated cells
[82]
model of colitis and TNF-α-induced HT29 treated cells
mRNA, ↑p-ERK, p-JNK, ↑pp38, ↑phosphorylation of the NF-κB protein and ↓PPAR-γ
submucosal edema, and ↑epithelium destruction
[82]
[82]
0, 12.5, 50, 100, 150, and 200 µM incubated for 24 h
↓p-ERK, ↓p-JNK, ↓p-p38, ↓phospho-NF-kB protein and ↑PPAR-γ in vivo
phosphorylation of the ↓PPAR-γ
phosphorylation of the ↓PPAR-γ
↓VCAM1 mRNA, ↓iNOS mRNA, and vivo and
visible fecal blood,
↓VCAM1 mRNA, ↓iNOS COX-2 mRNA in
↓Body weight, loose feces,
7.5, 15, 30 mg/kg/
↑ ↑ visible fecal blood,
↓VCAM1 mRNA, ↓iNOS mRNA, and ↓COX-2 mRNA in vivo and ↓IL-1β expression, ↓IL-18 expression, ↓phosphorylation of IKKα/β, ↓IκBα, ↓phosphorylation of the p65, ↓p65, ↓Caspase-1 activation and ↓NLRP3 inflammasome in vitro
VCAM1 mRNA, ↓iNOS COX-2 mRNA in
IL-1β expression, ↓IL-18 phosphorylation
Fraxinellone
↓Body weight, ↑diarrhea, ↑loose feces, ↑visible fecal blood, ↑mortality, ↑gross bleeding, ↑ulcerations, colon length, ↑DAI, ↑inflammatory cell infiltration at mucosa and submucosa, ↑crypts distortion, and ↓goblet cells
Fraxinellone
↑IL-1β, ↑IL-18, vivo and ↑IL-1β, ↑NO in vitro
ulcerations, ↑DAI, ↑inflammatory cell infiltration at mucosa and submucosa, ↑crypts distortion, and ↓goblet
Fraxinellone
visible fecal blood, ↑mortality, gross bleeding, ulcerations, colon length, ↑DAI, ↑inflammatory cell infiltration at mucosa and submucosa, ↑crypts distortion, and ↓goblet cells
gross bleeding, ulcerations, colon length, ↑DAI, ↑inflammatory cell infiltration at mucosa and submucosa, ↑crypts distortion, and ↓goblet cells
[83]
IL-1β expression, ↓IL-18 expression, phosphorylation of IKKα/β, IκBα, ↓phosphorylation of the p65,
day intraperitoneally/9 days in vivo and 10, 30 µM incubated for 24 h
and 10, 30 µM incubated for 24 h in vitro
IκBα, ↓phosphorylation of the p65, ↓p65, ↓Caspase-1 activation
DSS-induced C57BL/6J mice model of colitis and LPS-induced Human
↑IL-1β, ↑IL-18, TNF-α, ↑IL-6 in vivo and ↑IL-1β, IL-18 and ↑NO in vitro
C57BL/6J mice model of colitis and LPS-induced Human THP-1 treated cells
DSS-induced C57BL/6J mice model of colitis and LPS-induced Human THP-1 treated cells
7.5, 15, 30 mg/kg/ day intraperitoneally/9 days in vivo and 10, 30 µM incubated for 24 h in vitro
↓IL-1β expression, ↓IL-18 expression, phosphorylation of IKKα/β, IκBα,
LPS-induced Human THP-1 treated cells
day intraperitoneally/9 days in vivo and 10, 30 µM
↑IL-1β, ↑IL-18, ↑TNF-α, ↑IL-6 in vivo and ↑IL-1β, ↑IL-18 and ↑NO in vitro
↑IL-1β, ↑IL-18, TNF-α, ↑IL-6 in vivo and IL-18 and ↑NO in vitro
[83]
[83]
[83]
↓NLRP3 inflammasome in
phosphorylation of the p65, Caspase-1 activation
Caspase-1 activation NLRP3 inflammasome in
NLRP3 inflammasome in vitro
DSS-induced Sprague-Dawley rats model of colitis and LPS-induced RAW264.7 treated cells
Artesunate
mg/kg/day orally/5 days in vivo and 5, 10, and 20 µg/mL incubated for 24 h in vitro
Artesunate
10, 30, and 50 mg/kg/day orally/5 days in vivo and 5, 10, and 20 µg/mL
↑DAI, ↓hemoglobin, ↓colon length, and ↑cell destruction
10, 30, and 50 mg/kg/day orally/5 days in vivo and 5, 10, and 20 µg/mL incubated for 24 h in vitro
DSS-induced Sprague-Dawley rats model of colitis and LPS-induced RAW264.7 treated cells
IFN-γ, TLR4, ↑p-NF-κB, ↑p-p38, ↑Bax, ↑caspase-9 and Bcl-2
DSS-induced Sprague-Dawley rats model of colitis and
↑TNF- α, ↑IL-8, ↑IFN-γ, ↑TLR4, ↑p-NF-kB, ↑p-p38, ↑Bax, ↑caspase-9 and ↓Bcl-2
↓TLR4, ↓p-NF-κB, ↓p-p38, ↓Bax, ↓caspase-9 and ↑Bcl-2
10, 30, and 50 mg/kg/day orally/5 days in vivo and 5, 10, and ↓colon length, and ↑cell
Artesunate
↑DAI, ↓hemoglobin, ↓colon length, and ↑cell destruction
↓TLR4, ↓p-NF-kB, ↓p-p38, ↓Bax, ↓caspase-9 and ↑Bcl-2
[84]
DSS-induced Sprague-Dawley rats model of colitis and LPS-induced
[84]
Artesunate
↑DAI, ↓hemoglobin,
↑TNF- α, ↑IL-8, IFN-γ, ↑TLR4,
↓TLR4, ↓p-NF-κB, ↓p-p38,
IFN-γ, ↑TLR4, ↑p-NF-κB, ↑p-p38, ↑Bax,
↑Bax, ↑caspase-9 and ↓Bcl-2
[84]
destruction
↓Bax, ↓caspase-9 and ↑Bcl-2
- Metabolites 2023, 13, 96 22 of 47
THP-1 treated cells
infiltration at mucosa
↓p65, ↓Caspase-1 activation
in vitro
Table 1. Cont.
10, 30, and 50 mg/kg/day orally/5 days in
Related Molecular Mechanisms in Regulation of NF-kB in IBD
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
Sprague-Dawley rats model of colitis and LPS-induced
Ref.
Phytochemicals
↑DAI, hemoglobin,
↑TNF- α, ↑IL-8, IFN-γ, ↑TLR4,
↓TLR4, ↓p-NF-κB, ↓p-p38, ↓Bax, ↓caspase-9 and ↑Bcl-2
↑ destruction
↑caspase-9 and ↓Bcl-2
20 µg/mL
↓iNOS expression, ↓p–NF–κB P65 expression, ↓NF-kB-P65 nuclear translocation, ↓p38 phosphorylation, ↓JNK phosphorylation, ↓ERK phosphorylation and ↓NLRP3 expression in vitro and ↓NF-kB-P65, ↓p38 phosphorylation, ↓JNK phosphorylation and ↓ERK phosphorylation in vivo
18 of 44
18 of 44
↑NO, ↑iNOS expression, ↑p–NF–κB-P65 expression, ↑NF-kB P65 nuclear translocation, ↑p38 phosphorylation, ↑JNK phosphorylation, ↑ERK phosphorylation, ↑NLRP3 expression in vitro and ↑NF-kB P65, ↑TNF-α and ↑IL-6 in vivo
Colon length, ↓body
NO, iNOS expression, ↑p–
↓iNOS expression, ↓p–NF–κB P65 expression, ↓NF-κB-P65 nuclear translocation, ↓p38 phosphorylation, ↓JNK
Aesculetin DSS-induced C57BL/6J mice model of colitis and LPS-induced
20 mg/kg/day orally/ 7 days in vivo and 10, 25, 50 µM incubated for 4 h in vitro
↓Colon length, ↓body weight, ↑DAI, and ↑inflammatory cell infiltration
DSS-induced C57BL/6J mice model of colitis and LPS-induced RAW264.7 treated cells
20 mg/kg/day orally/7 days in vivo and 10, 25, 50
Aesculetin
NF–κB-P65 expression, ↑NF-κB P65 nuclear translocation, ↑p38 phosphorylation, ↑JNK
phosphorylation, ↑ERK phosphorylation, ↑NLRP3
phosphorylation, ↓ERK phosphorylation and ↓NLRP3 expression in vitro and ↓NF-κBP65, ↓p38 phosphorylation, ↓JNK phosphorylation and
weight, DAI, and ↑inflammatory cell
RAW264.7 treated cells
µM incubated for in vitro
phosphorylation, ↑ERK phosphorylation, ↑NLRP3
phosphorylation, ↓ERK phosphorylation and ↓NLRP3 expression in vitro and ↓NF-κBP65, ↓p38 phosphorylation,
[85]
RAW264.7 treated cells
µM incubated for in vitro
[85]
and ↑NF-κB IL-6 in vivo
and ↑NF-κB IL-6 in vivo
JNK phosphorylation and ERK phosphorylation in vivo
ERK phosphorylation in vivo
↑IL-1β, ↑CXCL1, ↑MIP-2, ↑MCP-1, ↑IL-1β mRNA, ↑CXCL1 mRNA, TNF-α mRNA, ↑IL-6 mRNA, ↑NOS2 expression, VEGF expression, ↑Ki67 expression, ↑NF-kB-p65 phosphorylation, ↑ICAM-1 mRNA, VCAM-1 mRNA and
↑Hemorrhage in the colonic lumen, ↓body
↑IL-1β, ↑CXCL1, ↑MIP-2, ↑MCP-1, ↑IL-1β mRNA, ↑CXCL1 mRNA, ↑TNF-α mRNA, ↑IL-6 mRNA, ↑NOS2 expression, ↑VEGF expression, ↑Ki67 expression, ↑NF-kB-p65 phosphorylation, ↑ICAM-1 mRNA, ↑VCAM-1 mRNA and ↑LFA-1 mRNA
MIP-2, ↑MCP-1, ↑IL-1β mRNA, ↑CXCL1 mRNA, TNF-α mRNA, ↑IL-6 mRNA, ↑NOS2 expression, VEGF expression, ↑Ki67 expression, ↑NF-kB-p65 phosphorylation, ↑ICAM-1 mRNA, VCAM-1 mRNA and ↑LFA-1 mRNA
↑Hemorrhage in the colonic
colonic lumen, ↓body weight, diarrhea with bloody stools, ↑DAI, ↑mucosal neutrophils infiltration, crypts,
Euphol
Euphol
Euphol
lumen, ↓body weight, ↑diarrhea with bloody
diarrhea with bloody stools, ↑DAI, ↑mucosal neutrophils infiltration, crypts, ↓ ↑mucosal
DSS and TNBSinduced CD1 mice model of colitis and LPS-induced
mg/kg twice a day orally for 3, 4 or 7 days in vivo and 1 and 10 µM
DSS and TNBS-induced CD1 mice model of colitis and LPS-induced BMDMs treated cells
3, 10, and 30 mg/kg twice a day orally for 3, 4 or 7 days in vivo and 1 and 10 µM incubated for 24 h in vitro
↓NOS2 expression, ↓VEGF expression and ↓p65 NF-kB activation
induced CD1 mice model of colitis and LPS-induced
orally for 3, 4 or 7 days in vivo and 1 and 10 µM
↓NOS2 expression, ↓VEGF expression and ↓p65 NF-kB activation
↓NOS2 expression, ↓VEGF expression and ↓p65 NF-kB activation
stools, ↑DAI, ↑mucosal neutrophils infiltration,
[86]
[86]
[86]
↓crypts, ↓goblet cells, ↑mucosal hyperemia, ↑mucosal necrosis
↑mucosal mucosal
mucosal
necrosis
MLCK mRNA, ↓MLCK
↓Akt, ↓MLCK mRNA, ↓MLCK protein and ↓NF-kB p65 protein expression in vitro and ↓MLCK, ↓NF-kB, ↓phosphatidylinositol 3-kinase (PI3K), ↓Akt, ↓NF-kB p65 protein expression, ↓iNOS expression and ↓COX-2 expression in vivo
MLCK mRNA, ↓MLCK
↑Akt, ↑MLCK mRNA,
colon weight-to-length intestinal permeability, ↑MPO, ↑TNF-α, ↑IL-1β, ↑IL-6, ↑NO, ↑PGE2, ↑iNOS expression, COX-2
NF-kB p65 protein expression in vitro and ↓MLCK, ↓NF-kB, phosphatidylinositol 3-kinase (PI3K), ↓Akt, ↓NF-kB p65 protein expression, ↓iNOS expression and ↓COX-2 expression in vivo
↑DAI, ↓body weight, ↑colon
↑Akt, ↑MLCK mRNA, ↑MLCK
colon weight-to-length Ratio, intestinal permeability, ↑MPO, ↑TNF-α, ↑IL-1β, ↑IL-6, ↑NO, ↑iNOS
protein and NF-kB p65 protein expression in vitro and ↓MLCK, ↓NF-kB, phosphatidylinositol
Nobiletin
↑Akt, ↑MLCK mRNA, ↑MLCK protein and ↑NF-kB p65 protein expression in vitro and ↑MLCK, ↑NF-kB, ↑PI3K, ↑Akt and ↑NF-kB p65 protein Expression in vivo
protein and ↑NF-kB p65 protein expression and ↑MLCK, ↑NF-kB, PI3K, Akt and ↑NF-
Nobiletin
Nobiletin
weight-to-length Ratio, ↑intestinal permeability, ↑MPO,
protein and NF-kB p65 protein expression and ↑MLCK, ↑NF-kB, ↑PI3K, Akt and ↑NF-
Sprague-Dawley rats model of colitis and LPS-induced Caco-2 treated cells
orally/7 days vivo and 0, 10, 20, 40, or 80 incubated for 0–36 h µM in
TNBS-induced Sprague-Dawley rats model of colitis and LPS-induced Caco-2 treated cells
20 and 40 mg/kg/day orally/7 days in vivo and 0, 10, 20, 40, or 80 incubated for 0–36 h µM in vitro
Sprague-Dawley rats model of colitis and LPS-induced Caco-2 treated cells
orally/7 days in vivo and 0, 10, 20, 40, or 80 incubated for 0–36 h µM in
[87]
[87]
3-kinase (PI3K), ↓Akt, ↓NF-kB p65 protein expression, ↓iNOS expression and ↓COX-2
↑TNF-α, ↑IL-1β, ↑IL-6, ↑NO, ↑PGE2, ↑iNOS expression, ↑COX-2 expression in vivo
kB p65 protein Expression in vivo
kB p65 protein Expression in vivo
COX-2 in vivo
in vivo
in vivo
Galangin
↑mucosal necrosis, ↓TLR4 mRNA, ↓NF-κB-p65
activity index; DMSO, dimethylsulfoxide; DNA, deoxyribonucleic acid; DNCB, dinitrochlorobenzene; DSS, dextran sulfate sodium; ERK, extracellu-
- Metabolites 2023, 13, 96 23 of 47
20 and 40
Sprague-Dawley rats model of colitis and LPS-induced Caco-2 treated cells
Table 1. Cont.
↑TNF-α, ↑IL-1β, ↑IL-6,
↑NF-kB, ↑PI3K, ↑Akt and ↑NF-
3-kinase (PI3K), ↓Akt, ↓NF-kB p65 protein expression, ↓iNOS expression and COX-2 expression in vivo
40, or 80 incubated for 0–36 h µM in
Related Molecular Mechanisms in Regulation of NF-kB in IBD
In Vivo/In Vitro Model(s)
Effective Dose(s)/ Concentration(s)
Related Clinical Symptoms of IBD
NF-kB-Related Dysregulation Indicators
Ref.
Phytochemicals
expression, COX-2 expression
vivo
↑Mucosal ulceration,
Galangin
Galangin
↑Mucosal ulceration, ↑mucosal necrosis, ↑inflammatory cell infiltration in the lamina propria and submucosa
mucosal necrosis, ↑inflammatory cell infiltration in the lamina propria and
NF-κB-p65 nuclear translocation, ↓TNF-α expression, and ↓IL-6 expression
↓TLR4 mRNA, ↓NF-kB-p65 nuclear translocation, ↓TNF-α expression, and ↓IL-6 expression
↑TLR4 mRNA, NF-κB p65 nuclear translocation, ↑TNF-α and ↑IL-6
↑TLR4 mRNA, ↑NF-kB p65 nuclear translocation, ↑TNF-α and ↑IL-6
40 mg/kg/day orally for 20 days
DSS-induced Swiss albino mice model of colitis
40 mg/kg/day orally for 20 days
albino mice model of colitis
[88]
[88]
, increase; ↓, decrease; AcOH, acetic acid; ALP, alkaline phosphatase; Akt, protein kinase b; AP-1, activating protein-1; ASC, apoptosis-associated speck-like protein; Bax, Bcl-2-associated X protein; Bcl-2, B-cell lymphoma 2 protein; BMDMs, bone-marrow-derived macrophages; Caco-2, human colorectal adenocarcinoma cells; CAS, Clinical activity score; CAT, catalase; CD1b, CD1b T cell surface glycoprotein; CD11b, integrin alpha M; CD3, cluster of differentiation 3, c-Fos, cellular proto-oncogene Fos; cl-caspase3, cleaved Caspase-3; CMDI, colon macroscopic damage index; C-MYC, cellular myelocytomatosis oncogene; COX-2, cyclooxygenase 2; CRP, c reactive protein; CXCL-1, chemokine (C-X-C motif) ligand 1; DAI, disease activity index; DMSO, dimethylsulfoxide; DNA, deoxyribonucleic acid; DNCB, dinitrochlorobenzene; DSS, dextran sulfate sodium; ERK, extracellular signal-regulated kinase; GRK2, G-protein-coupled receptor kinase 2; GSK3β, glycogen synthase kinase 3 beta; GSH, glutathione; GSH-Px, glutathione peroxidase; HCT-116, human colorectal carcinoma cell line; HFD, high-fat diet; HO-1, heme oxygenase-1; hPXR, human pregnane X receptor; HT-29, human colorectal adenocarcinoma cell line; ICAM-1, Intercellular Adhesion Molecule 1; ICR, Institute of Cancer Research; IEC-6, intestinal epithelioid cell 6; IFN-γ, interferon gama; IκBα, nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor alpha; IKK, multi-subunit IkB kinase; IKKβ, inhibitor of nuclear factor kappa-B kinase subunit beta; IL-1, interleukin 1; IL-1β, interleukin 1 beta; IL-2, interleukin 2; IL-4,
↑, increase; ↓, decrease; AcOH, acetic acid; ALP, alkaline phosphatase; Akt, protein kinase b; AP-1, activating protein-1; ASC, apoptosis-associated speck-like protein; Bax, Bcl-2-associated X protein; Bcl-2, B-cell lymphoma 2 protein; BMDMs, bone-marrow-derived macrophages; Caco-2, human colorectal adenocarcinoma cells; CAS, Clinical activity score; CAT, catalase; CD1b, CD1b T cell surface glycoprotein; CD11b, integrin alpha M; CD3, cluster of differentiation 3, c-Fos, cellular proto-oncogene Fos; cl-caspase3, cleaved Caspase-3; CMDI, colon macroscopic damage index; C-MYC, cellular myelocytomatosis oncogene; COX-2, cyclooxygenase 2; CRP, c reactive protein; CXCL-1, chemokine (C-X-C motif) ligand 1; DAI, disease activity index; DMSO, dimethylsulfoxide; DNA, deoxyribonucleic acid; DNCB, dinitrochlorobenzene; DSS, dextran sulfate sodium; ERK, extracellular signal-regulated kinase; GRK2, G-protein-coupled receptor kinase 2; GSK3β, glycogen synthase kinase 3 beta; GSH, glutathione; GSH-Px, glutathione peroxidase; HCT-116, human colorectal carcinoma cell line; HFD, high-fat diet; HO-1, heme oxygenase-1; hPXR, human pregnane X receptor; HT-29, human colorectal adenocarcinoma cell line; ICAM-1, Intercellular Adhesion Molecule 1; ICR, Institute of Cancer Research; IEC-6, intestinal epithelioid cell 6; IFN-γ, interferon gama; IκBα, nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor alpha; IKK, multi-subunit IkB kinase; IKKβ, inhibitor of nuclear factor kappa-B kinase subunit beta; IL-1, interleukin 1; IL-1β, interleukin 1 beta; IL-2, interleukin 2; IL-4, interleukin 5; IL-6, interleukin 6; IL-10, interleukin 10; IL-12, interleukin 12; IL-15, interleukin 15; IL-17, interleukin 17; IL-18, interleukin 18; IL-27, interleukin 27; iNOS. inducible nitric oxide synthase; IRAK1, interleukin 1 receptor associated kinase 1; JAM-A, junctional adhesion molecule A; JAK1, Janus kinase 1; JAK2, Janus kinase 2; JNK, c-Jun N-terminal kinases; Ki67, antigen KI-67; KM, Kunming Mouse; LEF-1, lymphoid enhancer binding factor 1; LFA-1, lymphocyte function-associated antigen 1; LDH, lactate dehydrogenase; LPO, lactoperoxidase; LPS, lipopolysaccharide; LRP6, low-density lipoprotein receptor-related protein 6; LTB4, leukotriene B4; Ly6G, lymphocyte antigen 6 complex locus G; MAPK, mitogen-activated protein kinase; MAPK1, mitogen-activated protein kinase 1; MCP-1, monocyte chemoattractant protein-1; MDA, malonaldehyde; MD2, myeloid differentiation protein 2; MDSC, myeloid-derived suppressor cells; MIP-1α, macrophage inflammatory protein 1 α; MIP-2, macrophage inflammatory protein-2; MMP-9, matrix metalloproteinase-9; MLCK, myosin light-chain kinase; MPO, myeloperoxidase; mRNA, messenger RNA; Muc2, Mucin 2, oligomeric mucus/gel-forming; MyD88, MYD88 innate immune signal transduction adaptor; NADPH, nicotinamide adenine dinucleotide phosphate; NF-kB, nuclear factor kappa b; NF-kBp65, NF-kB classical signaling pathway protein; NLRP3, NLR [nucleotide-binding domain leucine-rich repeat] family pyrin domain containing 3; NO, nitric oxide; NOS2, nitric oxide synthase 2 (inducible); Nrf2, nuclear factor erythroid 2–related factor 2; OS, oxidative stress; p-, phosphorylation; p-AKT, phosphorylated protein kinase B; PGD2/PGE2, prostaglandin D2; PGE2, prostaglandin E2; PGJ2, prostaglandin J2; PI3K, phosphoinositide 3-kinase; pNF-kB, phospho-NF-kB p65; PPARγ, peroxisome proliferator- activated receptor gamma; ROS, reactive oxygen species; RSV, resveratrol; SAPK, stress-activated protein kinases; STAMP2, six transmembrane protein of prostate 2; STAT1, signal Transducer And Activator Of Transcription 1; SIRT1, NAD-dependent deacetylase sirtuin-1; SIgA, immunoglobulin A; SIGIRR, single Ig IL-1-related receptor; SOD, superoxide dismutase; STAT3, signal transducer and activator of transcription-3; TAK1, transforming growth factor-β-activated kinase 1; TBARS, thiobarbituric acid reactive substances; TCF-4, transcription factor 4; TDNPs, turmeric-derived nanoparticles; TGF-β, transforming growth factor beta; THP-1, human leukemia monocytic cell line; TLR4, toll-like receptor 4; TLR4-NF-kB-NLRP3, toll-like receptor 4-nuclear factor kappa b-NLR family pyrin domain containing 3; TLR9, toll-like receptor 9; TNBS, 2,4,6-trinitrobenzene sulfonic acid; TNF-α, tumor factor necrosis alfa; TRIF, TIR-domain-containing adapter-inducing interferon-β; VCAM-1, vascular cell adhesion protein 1; VEGF, vascular endothelial growth factor; Wnt-1, proto-oncogene Wnt-1; ZO-1, zonula occludens-1; ZO-2, zonula occludens-2.
3. Results
This manuscript comprised data from 56 different phytocompounds and their derived secondary metabolites on in vitro and in vivo models of IBD. One study was only in vitro, 46 were in vivo, and 25 were both in vitro and in vivo studies. The included in vitro studies used many different cell models of IBD (such as RPMI 1640 treated Colon-26 cells, DMEM-treated Caco 2BBE, RAW 264.7 treated cells, LPS-treated human colorectal adenocarcinoma cells (Caco-2) cells, TNF-α-stimulated human colorectal carcinoma cell line (HCT-116) and human colorectal adenocarcinoma cell line (HT-29), and 2,4,6trinitrobenzene sulfonic acid (TNBS) -induced C3H/HeN). In addition, the in vivo models were a dextran sulfate sodium (DSS)-induced mice model of colitis, NFκB-RE-Luc transgenic models of colitis, TNBS-induced mice model of colitis, TNBS-induced Wistar Hannover rats model of colitis, TNBS-induced Wistar Hannover rats model of colitis, and Acetic acid-induced mice models of colitis. The included phytochemicals were curcumin, resveratrol, 3-(4-hydroxyphenyl)-propionic acid, sesamol, kaempferol, astragalin, pinocembrin, oxyberberine (bacterial metabolite), berberine hydrochloride, berberine, eriodictyol, betulin, naringin, 5-hydroxy-4-methoxycanthin-6-one, geniposide, sesamin, taxifolin, isobavachalcone, d-pinitol, paeoniflorin-6 -O-benzene sulfonate, thymol, tricin, aesculin, ginsenoside Rk3, lancemaside A, tetramethylpyrazine, daurisoline, tetrandrine, diosgenin, mangiferin, tryptanthrin, l-theanine, koreanaside A, 6-gingerol, lycopene, α-mangostin, ophiopogonin D, alantolactone, sinomenine, convallatoxin, fisetin, genipin, piperine, ligustilide, evodiamine, chrysin, wogonoside, oxymatrine, epicatechin, thymoquinone, fraxinellone, artesunate, aesculetin, euphol, nobiletin, and galangin.
4. Discussion
- 4.1. Physiopathology of Ulcerative Colitis
Pathologically, UC involves the intestinal epithelial barrier, gut commensal microbiota, and the immune system. The disruption of the tight junctions and the mucus film covering the lumen intestinal epithelial barrier layer promotes a state of increased permeability of the intestinal epithelium to luminal agents. At this point, commensal bacteria start to invade the other layers of the bowel’s tissues, and macrophages and dendritic cells, cells of the innate immune system, begin to recognize the bacterial antigen of the commensals through the utilization of pattern recognition receptors, such as the called Toll-like receptors (TLRs). Thus, these cells get over a tolerogenic status to an active phenotype in which activating pro-inflammatory pathways stimulates the transcription of multiple pro-inflammatory genes to promote the production and secretion of many pro-inflammatory cytokines like TNF-α, IL-12, IL-23, IL-1B, and IL-6 [89–91].
After the initial processing of the commensal bacterial antigens, the stimulated dendritic cells and macrophages start to present the antigens to naïve cluster of differentiation (CD) 4 (CD4) T cells, which are the main ones responsible for promoting T cells differentiation into Th (T helper) 2 effector cells, which are characterized by the production of a specific interleukin, which is the IL-4. Although naïve CD4 T cells are the primary regulators of the immune response, many other immune cells are involved pathogenically in UC development. For example, the natural killer (NK) cells are the main source of IL-13 during the disease process. This interleukin has been associated with the massive disruption of the intestinal lumen epithelial cell barrier. The inflamed intestine starts to express more and more adhesive molecules to leukocytes, leading to the increased entry of T cells into the intestinal wall lamina propria. Moreover, the intestinal cells have up-regulation of pro-inflammatory chemokines like C-X-C motif chemokine ligand (CXCL) 1 (CXCL1), CXCL3, and CXCL8, which only perpetuates the inflammatory cycle of colitis [89–92].
- 4.2. Physiopathology of Crohn’s Disease
- 4.3. NF-kB and Its Related Molecular Insights into Inflammation
NF-kB, or nuclear factor of the κ-chain in B-cells, is an inducible transcription factor that regulates many different genes involved in developing and regulating inflammatory and immunomodulated processes. NF-kB is composed of five structurally-related members, which are the NF-kB1 (p50), NF-kB2 (p52), ReIA (p65), ReIB, and c-ReI. These entities mediate the transcription of mainly inflammatory factors by binding to specific deoxyribonucleic acid (DNA) elements named kB enhancers and forming various hetero- and homo-dimers. NF-kB-related proteins are sequestered in the induced cells’ cytoplasm by a family of inhibitory proteins called IkB, characterized by the presence of ankyrin repeats. The most studied member is the nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor alpha (IkBα). Generally, the NF-kB’s vital activators are pro-inflammatory cytokines (IL-1β, TNF-α, IL-12, IL-33, IL-17, granulocyte macrophage-colony stimulating factor (GM-CSF), and lymphotoxin-β), bacterial antigens (lipopolysaccharides (LPS) such as flagellin, CpG-DNA, and enterotoxins), viral agents (viral proteins), many receptor ligands (CD40L, B-cell activating factor (BAFF), Fas ligand (FasL), hepatocyte growth factor (HGF), bone morphogenetic proteins (BMP) 2 (BMP-2), BMP-4, and TNF-related apoptosis inducing ligand (TRAIL)), cell lysis products (damage-associated molecular patterns (DAMPs), high mobility group box 1 (HMGB1), extracellular RNA molecules, and extracellular DNA molecules), eukaryotic parasites (Candida albicans and Leishmania), physiological stresses (endoplasmic reticulum (ER) stress, OS, acidic pH, hyperglycemia), physical stress (ionizing radiation and ultraviolet (UV)-light), and presence of modified proteins and particles (advanced glycation end products (AGEs), oxidized low-density lipoprotein cholesterol (LDL), and amyloid protein fragments) [98–100].
Molecularly, NF-kB activation involves two major signaling pathways, the canonical and the noncanonical (or alternative). Canonically, the NF-kB pathways are activated through several stimuli such as ligands of various pro-inflammatory cytokines receptors, T cell receptor (TCR), B cell receptor (BCR), TNF receptor (TNFR) superfamily member, and pattern-recognition receptor (PRRs). To enter the canonical pathway, NF-kB starts with the inducible degradation of IkBα, which is triggered by its site-specific phosphorylation by a multi-subunit IkB kinase (IKK) complex. The IKK complex can be activated by proinflammatory cytokines, growth factors, microbial components, mitogens, and stress agents and is formed of two catalytic subunits, which are the IKKα and IKKβ, additionally to a regulatory subunit called NF-kB essential modulator (NEMO) or IKKγ. Upon activation, the
IKK complex phosphorylates IKKα in two N-terminal serines. At this point, IKK triggers the ubiquitin-dependent IKKα in a proteasome that translocates rapidly and transiently the canonical NF-kB members from the cell’s nucleus. These canonical members are the p50/ReIA and the p50/c-ReI dimers [98,101–103].
In turn, the non-canonical (alternative) NF-kB activation is more selective. This pathway responds only to a specific stimuli group, including the ligands of a subset of the TNFR superfamily members (Lymphotoxin beta receptor [LTβR], tumor necrosis factor receptor superfamily member 13C (BAFFR), receptor activator of nuclear factor kappa-B (RANK), and CD40). Different from the canonical, the non-canonical pathway does not involve the degradation of the IkBα molecules but relies on processing the NF-kB2 (p100) precursor, an NF-kB-related protein. The processing of the p100 consists of the degradation of its C-terminal IkB-like structure, which results in the generation of a mature NF-kB2 p52 and the nuclear translocation of the NF-kB complex p52/ReIB. This degradation process is mediated by a central signaling molecule called NF-kB-inducing kinase (NIK), which functionally activates the IKKα and cooperates with its mediation on p100 phosphorylation. At this point, the phosphorylation of the p100 induces the p100 ubiquitination and further processing into the aforementioned NF-kB2 p52 and its nuclear translocation [98,104–106].
Theoretically, the canonical activation of the NF-kB is involved mainly in the development and progression of immunological responses. On the other hand, the non-canonical NF-kB activation is more involved in the supplementation signaling axis that cooperates with the canonical activation of the pathway in regulating specific adaptative functions of the adaptive immune system. To target inflammation, NF-kB not only exerts increases in the production of pro-inflammatory cytokines, adhesion molecules, and chemokines but also regulates the immune cells’ differentiation, apoptosis, morphogenesis, and proliferation. Dendritic cells are stimulated to maturation. T cells are stimulated to inflammatorymediated differentiation and activation and to form memory T cells principally through the actions of IL-12, IL-23, and RORγt, which are factors directly related to the NF-kB signaling. Macrophages are stimulated to massive production of pro-inflammatory cytokines and chemokines, in addition, to polarizing into M1. Neutrophils are stimulated into anti-apoptosis states and massive recruitment into inflammation sites [98,101,105,107].
In other words, NF-kB induces inflammatory cells survival due to the production of anti-apoptotic factors (such as Fas, B-cell leukemia/lymphoma 2 protein (Bcl-2), caspases, BFL-1, survivin, and cellular FLICE (FADD-like IL-1β-converting enzyme)-inhibitory protein (c-FLIP)), cell cycle regulators (like Bcl-2-like protein 1 (Bcl-2L1), Plasminogen activator inhibitor-2 (PAI2), and cyclin) and inflammatory cells proliferation due to the production of cells cycle regulators (IL-1, TNF-α, IL-2, IL-8, and IL-12). NF-kB also augments inflammatory cells’ adhesion and invasion into inflammatory sites due to augmentation of adhesion molecules production, such as intercellular adhesion molecule 1 (ICAM-1), epithelial cell adhesion molecule-1 (ECAM-1), vascular cell adhesion molecule 1 (VCAM-1), MMPs, and selectin. NF-kB-related pro-inflammatory cytokines and chemokines (monocyte chemoattractant protein-1 (MCP-1), IL-18, CXCL1, CXCL10, macrophage inflammatory proteins (MIP) 2 (MIP-2), and regulated upon activation, normal T cell expressed and secreted (RANTES)) also work on inducing angiogenesis in areas near inflammatory sites that help modulate the inflammatory response [98,101,104,105].
Biologically, NF-kB has a major function corresponding to changing cellular programs across different types of stress and harmful situations. During NF-kB activation and signaling, different cells are stimulated to cope with the threat and activate their defense mechanisms to profoundly eliminate the endangering and escape death with the final aim of returning to the original physiological state in which all cells of the body are organized into homeostasis. That is why NF-kB targets to upregulate or induce a variety of genes to elucidate its responses [101]. Due to its inflammatory nature and role in the body’s defense, NF-kB is implicated in the physiopathology and progression of several inflammatory diseases, such as IBD [98,108,109].
NF-kB triggers inflammatory diseases mainly by contributing to the activation and regulation of different inflammasomes. As a group of intracellular multi-protein complexes, inflammasomes assemble activating caspases principally in response to the presence of DAMPs and pathogen-associated molecular patterns (PAMPs). Inflammasomes are integrative parts of innate immunity and regulate the gut microbiota composition (a primary reason why NF-kB is involved with IBD). They are composed of a ligand-sensing receptor (usually a member of the nucleotide-binding domain leucine-rich repeat (NLR) family NLRP1, NLRP3, NLRC4, or the absent in melanoma 2 [AIM2]), one adaptor protein (potentially the apoptosis-associated speck-like protein containing CARD), and a pro-caspase (generally, the pro-caspase 1). Upon appropriate stimulation, the inflammasome receptors start to oligomerize and recruit pro-caspases 1 via apoptosis-associated Speck-like protein (ASC). At this point, pro-caspases 1 are converted into active caspases 1, and cleave the pro-IL-1β and pro-IL-18 into their mature forms. Then IL-1β and IL-18 are secreted as pro-inflammatory cytokines, and most inflammasome-derived inflammatory processes begin. The NLRP3 is the most extensive inflammasome ever studied, and it is composed of the NLRP3, ASC, and never in mitosis gene A (NIMA)-related kinase 7 (NEK7, an essential regulatory protein), and pro-caspases 1. When dysbalanced and disrupted, NF-kB super-activates inflammasomes, and an inflammatory disease arises [98,108,110].
- 4.4. NF-kB and Its Implications on IBD
NF-kB is implicated in the pathogenesis of IBD insofar as this transcription factor turned out to be the primary regulatory component of the inflammatory burden during the intestinal inflammation during UC and CD. The NF-kB signaling is not usually activated in regular patients but is activated in the inflamed bowel of patients affected by IBD. Immunofluorescence staining assays demonstrated that NF-kB activation in the intestinal lumen is predominant in the intestinal macrophages and epithelial cells of the intestinal lumen mucosa and, interestingly, the higher the number of cells with activated NF-kB stain, the worse the severity of the intestinal inflammation. In addition to macrophages and epithelial cells, other parts of the intestinal mucosa can express the NF-kB signaling pathways, as in the lamina propria fibroblasts, which are also assumed to play pro-inflammatory stimuli during IBD [111,112].
Especially in intestinal macrophages, NF-kB expression is accompanied by increased production of pro-inflammatory cytokines, such as TNF-α, IL-1, and IL-6, due to the actions of NF-kB in expanding the capacity of these cells to produce inflammatory mediators. In other intestinal cells, NF-kB also regulates the expression of IL-12 and IL-23. Predominantly, the NF-kB-derived pro-inflammatory cytokines are responsible for two mechanisms involved in the development of IBD lesions. Firstly, the produced pro-inflammatory cytokines directly mediate mucosal tissue damage by principally up-regulation of matrix metalloproteinases production and secretion in the intestinal lumen. Secondly, NF-kB-derived pro-inflammatory cytokines also mediate the stimulation, activation, and differentiation of immunological cells derived from lamina propria of the bowel mucosa, resulting in chronic inflammation and, therefore, in the perpetuation of mucosal damage. In this case, NF-kB acts crucially during principally CD development, in which Th1 cells are stimulated due to the major production of TNF-α and IL-23, and the TNF-α potentializes the proper NF-kB in a kind of positive feedback [111].
Besides the roles of NF-kB in macrophage-derived inflammatory patterns, which are well established, the actions of the signaling pathway in epithelial cells during the setting of IBD are still controversial. IL-6 and TNF-α majorly activate the epithelial cells at the beginning of the inflammatory process, leading to IBD. NF-kB is then activated, and this activation can be demonstrated by the increased intestinal lumen epithelial expression of a different cluster of adhesion molecules, such as the ICAM-1, which helps in the leukocyte recruitment to the inflammatory intestinal sites, such as in the case of neutrophils granulocytes. NF-kB also affects colonic lamina propria fibroblasts via activation of T-cellderived CD40 ligand (CD40L). The CD40L interacts with the CD40 receptor expressed
by the colonic fibroblasts on their surface and, thereby, induces the activation of the proinflammatory NF-kB in these cells, stimulating the increase in the expression of many different pro-inflammatory cytokines, such as IL-6, IL-8, and chemoattractant molecules, such as the monocyte chemoattractant protein [111].
Mainly in UC, recent research has revealed that immunological and inflammatory mechanisms are not unique in exerting the biological structural changes of colonic intestinal epithelial cells during the ulcerative phases of the disease. Ferroptosis is a condition of the iron-dependent type of nonapoptotic cell death that has recently emerged as a regulatory mechanism of the necrosis process observed in UC patients. Iron accumulation plays a critical role in cell death due to the increased accumulation of Reactive Oxygen Species (ROS) and the derived lipid peroxidation. Morphologically, ferroptosis appears
- as shrunken mitochondria, condensed mitochondrial membranes, and reductions in the mitochondrial crista. Additionally, ferroptosis also is related to a downregulation of the antioxidant systems, such as glutathione peroxidase (GPX) 4 (GPX4), which corresponds to a potent scavenger of reactive lipid molecules, and of the up-regulation of prostaglandinendoperoxide synthase 2 (PTGS2). Many authors suggested that a specific subunit of the NF-kB, the Bp65 subunit (NF-kBp65), plays a role in ferroptosis inhibition insofar as the deletion of intestinal epithelial cells NF-kBp65 was an up-regulator of ferroptosis that can exacerbate intestinal inflammation. There are suggestions that phosphorylated NF-kBp65 significantly inhibits endoplasmic reticulum stress signaling in the intestinal cells directly by binding a factor called eukaryotic initiation factor 2α. In this case, the ferroptosis process can be inhibited or tremendously decelerated [113].
Considering that upregulated NF-kB signaling pathway plays a critical role in IBD progression, Han et al. [114] hypothesized that different NF-kB activation levels influence CD’s clinical manifestations during the patient’s disease course. The results showed that higher NF-kB activation showed higher frequencies of ileocolonic involvement and lower frequencies of perianal lesions compared to patients with low NF-kB activation. Furthermore, higher activation demonstrated an association with higher histological scores. Lower NF-kB activity did not significantly affect disease progression course or outcome, principally after surgical treatment procedures.
Another important form of cell death that compromises the mucosal barrier integrity during IBD is pyroptosis; a caspase-1-dependent programmed cell death. This phenomenon features the gasdermin D (GSDMD) cleavage and causes the translocation and plasma membrane rupture of the mucosal cells, resulting in the massive release of pro-inflammatory cytokines to activate immunological mediators like IL-1β and IL-18. During IBD development, principally IL-1β works to alter the intestinal epithelial tight junctions, which results in increased intestinal permeability. Moreover, IL-18 has been shown to contribute to the decrease in the gut mucosal barrier integrity, causing inflammation and amplifying tissue damage. Mechanistically, pyroptosis can depend on a highly glycosylated transmembrane protein called CD147 to happen. CD147 induces pyroptosis in intestinal epithelial cells by enhancing the expression of IL-1β and IL-18, which activate inflammasomes, including caspase-1 and GSDMD, aggravating inflammatory reactions and events. It is known that CD147 stimulates the phosphorylation of NF-kBp65 in intestinal epithelial cells. Thus, inhibition of the CD147 molecule can correspond to a novel therapeutic strategy to combat IBD [112,115].
There is a long list of studies regarding the inhibition of NF-kB as a powerful treatment strategy for IBD. Qiu et al. [116] found that maresin 1, a specialized resolving mediator of inflammation derived from macrophages, has a powerful anti-NF-kB inhibition effect through toll-like receptor 4 (TLR4), which alleviated UC in a rat model of dextran sulfate sodiuminduce disease. Wang et al. [117] found by studying a rat model of acetic acid-induced UC that a methane-rich saline preparation could effectively promote an anti-inflammatory response against the disease by principally mediating the blockage of the NF-kB through TLR4 again. Tong et al. [118] found that milk alleviates colitis by regulating Treg cells and inhibiting TLR4/ NF-kB signaling pathways, restoring immunity balance and reshaping
gut microbiota. Chen et al. [119] found that sodium butyrate inhibits inflammation in a Trinitrobenzenesulfonic Acid (TNBS)-induced IBD mice model and maintains epithelium barrier integrity through activation of G protein coupled receptor 109A (GPR109A) and inhibition of the protein kinase B (Akt) and NF-kBp65 signaling pathways.
- 4.5. Phytochemicals That Influence the NF-kB Signaling during IBD: An Overview
As shown in the following items of this section, many phytochemicals have a critical role in regulating inflammatory processes observed in IBD. Table 1 summarizes some of these molecules and their actions
- 4.5.1. Curcumin
Curcuma longa (CL) is a plant belonging to Zingiberaceae, also known as turmeric. Widely used as a spice in food preparation, mainly in India, CL is grown in Southeast Asia, China, and Latin America. Polyphenolic bioactive compounds, called curcuminoids (CCMs), such as curcumin (CUR), demethoxycurcumin (DMC), and bisdemethoxycurcumin (BMC), were identified from this plant. Curcumin has therapeutic potential for anti-inflammatory, anti-diabetic, anti-cancer, and anti-aging effects [120]. In addition, studies have revealed that this compound can act therapeutically against chronic pulmonary, cardiovascular, neurological, neoplastic, psychological, and metabolic diseases [121]. CUR plays anti-inflammatory and immunomodulatory roles in the pathogenesis and progression of IBD. Studies showed that it stimulates the differentiation of intestinal Treg cells, and in experimental mouse models, these Tregs prevented the development of colitis. Other research indicated that pretreatment with CUR suppressed LPS-induced NF-kB-p65 translocation and mitogen-activated protein kinases (MAPK) phosphorylation in dendritic cells of colitis mice models, reducing inflammation [122].
Liu et al. [17] evaluated the effects of turmeric-derived nanoparticles containing high levels of curcumin in DSS-induced FVB/NJ female mice and NFκB-RE-Luc transgenic mice model of colitis, in addition to RPMI 1640-treated Colon-26 cells, DMEM-treated Caco2BBE and RAW 264.7 cells. The results showed that NF-kB translocation was inhibited in vitro and in vivo. In addition, the treatment decreased Myeloperoxidase (MPO) levels and mRNA expression of pro-inflammatory cytokines such as TNF-α, IL-1β, and IL-6.
Altinel et al. [18] aimed to investigate the roles of anal and oral CUR against NF-kB activity using a mouse model of TNBS-induced colitis. Compared to the control group, colonic levels of NF-kB had a more significant reduction with oral CUR. Furthermore, the expression of platelet-derived growth factor (PDGF) and TNF-α also lower in the oral CUR group when analyzed by biochemistry.
An in vivo study by Kao et al. [19] analyzed the effects of CUR on NF-kB deactivation in a murine model of DSS-induced colitis. The results showed that CUR could inhibit IKKβ activity by S-nitrosylation in colonic cells, a nitric oxide (NO) dependent modifier
- at the cysteine residue that regulates inducible nitric oxide synthase (iNOS) activity. In addition, CUR was able to sequentially inhibit the phosphorylation of IkB, actions that culminated in the inhibition of NF-kB in the intestine. CUR also repressed the production of pro-inflammatory cytokines such as TNF-α, IL-1β, and IL-6.
An in vivo study performed by Zeng et al. [20] with a murine model of TNBS-induced colitis evaluated the protective effects of CUR against the production of inflammatory factors in the intestine of the animals. The CUR-treated group showed a reduction in NF-kB and IL-27 mRNA expression compared to the untreated group. In addition, there was a significant reduction in NF-kB-p65 protein expression in the group that used CUR.
According to Lubbad et al. [21] study, using CUR in an in vivo model of TNBS-induced colitis resulted in decreased colonic activation of NF-kB. In addition, CUR decreased colonic concentrations of oxidative enzymes such as Malondialdehyde (MDA) and MPO.
Larmonier et al. [22] indicated that there was no concentration of CUR that modified NF-kB activation in the subepithelial tissue of IL-10−/−/NF-kBEGFP transgenic mice.
However, 0.1% CUR demonstrated protective effects against colitis and decreased NF-kB activation in colonic epithelial cells only in IL-10 / mice.
Venkataranganna et al. [23] used NCB-02, a CL extract with high concentrations of CUR, to assess its benefits on dinitrochlorobenzene (DNCB) -induced UC in rats. Results showed that NCB-02 inhibited the expression of NF-kB, iNOS, and MPO in treated UC mice. There were also improvements in colon weight and size and reduced damage to the gut structure.
Zhang et al. [24] induced TNBS colitis in an in vivo model to assess the effects of CUR on NF-kB deactivation. The animals treated with CUR had much smaller and more superficial ulcers than the untreated group. Furthermore, most of these ulcers were in an advanced state of regeneration. The group treated with CUR still showed a decrease in intestinal epithelial necrosis and infiltration of inflammatory cells in the submucosa and lamina propria of the intestine. Peroxisome proliferator-activated receptor γ (PPARγ) messenger RNA (mRNA) was increased in the colonic mucosa in the CUR-treated group, and CUR also increased the concentration of Prostaglandin E2 (PGE2) and 15-deoxydelta12,14-prostaglandin J2 (15d-PGJ2). Finally, there were reductions in mRNA expression of IL-1β, TNF-α, and IFN-γ, cyclooxygenase-2 (COX-2).
The experimental study by Jian et al. [25] in a rat model of TNBS-induced colitis evaluated the DNA binding activity of NF-kB. Rats were treated with CUR, stimulating IkB degradation and inhibiting IL-1β mRNA expression.
Ukil et al. [27] evaluated the effects of CUR on TNBS-induced colitis in mice and showed that sick animals presented an increase in NF-kB DNA binding activity in nuclear extracts from inflamed colonic tissue. CUR inhibited the binding of the NF-kB factor to its nuclear targets. Such CUR action was due to the inhibition of NF-kB mobility. It was also observed that the pre-treatment group with CUR showed a greater formation of Th2 response by suppressing IFN-y and IL-12 p40 mRNA and reducing the expression of iNOS mRNA. Serine protease activity, as well as neutrophilic infiltration and accumulation of MPO, MDA, and NO were also prevented by pretreatment with CUR.
Using a model of DNB-induced colitis in mice, Salh et al. [26] demonstrated that CUR leads to a reduction in NF-kB DNA binding activity and reduces MPO action and neutrophilic infiltration, as well as IL-1β mRNA expression.
Sugimoto et al. [28] evaluated the effects of CUR on TNBS-induced colonic inflammation in mice and showed a decrease in p65 expression and inhibited IkB degradation and NF-kB translocation in the nucleus of intestinal epithelial cells. Furthermore, CUR inhibited IkB degradation in colonic macrophages and reduced colonic inflammation by decreasing the infiltration of CD4 T cells into the lamina propria and the expression of pro-inflammatory cytokine genes in the colonic mucosa.
Figure 2 shows the regulation of NF-kB pathways with the use of curcumin. 4.5.2. Resveratrol
Resveratrol (RSV) is a phenolic compound mainly produced by plants in response to environmental stress. Primarily, RSV is considered an anti-inflammatory phytochemical insofar as it inhibits many pro-inflammatory pathways, including the NF-kB, and exerts antioxidant, anti-aging, and cardiovascular protection effects. This phenol has been encountered in more than 72 different plant species until now and exists in two geometric isomers, which are the trans- and cis-, and two glucosides, which are the trans- and cispiceids [123,124].
In a mice model of TNBS-induced colitis, Lu et al. [29] showed that RSV supplementation leads to NF-kB downregulation through decreases in pNF-kB, TNF-α mRNA, TNF-α, transforming growth factor beta (TGF-β) mRNA, and TGF-β. RSV also induced visceral hyperalgesia, an effect derived from the inhibition of spinal NF-kB signaling, which reduced the production of pro-inflammatory cytokines.
MetabolitesMetabolites2023, 132023, 96 31 of 47
Figure 2. NF-kB regulation with the use of curcumin. ↑, increase; ARE, antioxidant response elements; ASC, apoptosis-associated Speck-like protein; CAT, catalase; HO-1, heme oxygenase 1; GPX, glutathione; GST, glutathione S-transferases; IBD, inflammatory bowel diseases; Keap1, Kelch-like ECH-associated protein 1; MAPK, mitogen-activated protein kinases; NF-kB, nuclear factor-Kb; NLRP3, NLR (/nucleotide-binding leucine-rich repeat receptor) family pyrin domain containing 3; NQO1, NAD(P)H dehydrogenase (quinone) 1; Nrf2, nuclear factor erythroid 2–related factor 2; SOD1, superoxide dismutase 1; TAK1, transforming growth factor-β-activated kinase 1; TRAF6, tumor necrosis factor receptor (TNFR)-associated factor 6.
- 4.5.2. Resveratrol
- 4.5.3. 3-(4-Hydroxyphenyl)-propionic Acid
Zhang et al. [34] showed that in vitro and also in vivo, the use of 3-(4-hydroxyphenyl)propionic acid, which is a microbial metabolite of RSV and a bioactive compound richly found in Aloe africana, was able to reduce the expression of NF-kB-related activation proteins and MAPK. After the treatment, the models presented elevated inflammation and OS, markedly due to an elevated expression of NF-kB and MAPK.
cell apoptosis.
- Figure 3. Main regulatory effects of RSV on NF-kB modulation during IBD. IL-6, interleukin 6; IL-10, interleukin 10; IL-12, interleukin 12; IL-17, interleukin 17; MHC, major histocompatibility complex; MPO, myeloperoxidase; NF-kB, nuclear factor-kB NF-kB; NO, nitric oxide; ROS, reactive oxygen species; TCR, T-cell receptor; Th0, T helper 0; Th1, T helper 1; Th2, T helper 2; Th17, T helper 17; TLR, Toll-like receptor; TNF-α, tumor factor necrosis alfa; Treg, regulatory T cell.
- 4.5.4. Sesamol
- 4.5.5. Kaempferol
- 4.5.6. Astragalin
Figure 3. Main regulatory effects of RSV on NF-kB modulation during IBD. IL-6, interleukin 6; IL10, interleukin 10; IL-12, interleukin 12; IL-17, interleukin 17; MHC, major histocompatibility complex; MPO, myeloperoxidase; NF-kB, nuclear factor-kB NF-kB; NO, nitric oxide; ROS, reactive oxygen species; TCR, T-cell receptor; Th0, T helper 0; Th1, T helper 1; Th2, T helper 2; Th17, T helper 17; TLR, Toll-like receptor; TNF-α, tumor factor necrosis alfa; Treg, regulatory T cell.
Peng et al. [37] conducted a study with a DSS-induced C57BL/6 mice model of colitis to study the effects of astragalin, a bioactive compound of Moringa oleifera, Cassia alata, and Rosa agrestis, against the NF-kB activation in this model of IBD. The control mice presented increased DAI scores, intestinal mucosal injury, inflammatory cell infiltration,
and decreased colon length. After the treatment, the mice demonstrated downregulation of all pro-inflammatory cytokines mRNA, TLR4 mRNA and p-IκBα, p-IKKα/β, and p-p65 expressions, as well as increased expressions of ZO-1 mRNA, occludin mRNA, and Mucin 2, oligomeric mucus/gel-forming (Muc2) mRNA.
Han et al. [38] also studied astragalin treatment against IBD models. These authors used TNF-α -stimulated HCT-116 and HT-29 human colonic epithelial cells in vitro and DSS-induced C57BL/6 mice model of colitis in vivo and found that after the treatment, there was a decreased cell proliferation, TNF-α mRNA, IL-8 mRNA, IL-6 mRNA, IκBα phosphorylation, and NF-kB-DNA binding in vitro, as well as decreased TNF-α mRNA, IL-8 mRNA, IL-6 mRNA, and IκBα phosphorylation in vivo.
- 4.5.7. Pinocembrin
- 4.5.8. Oxyberberine
- 4.5.9. Berberine Hydrochloride
- 4.5.10. Berberine
Lee et al. [42] evaluated the effects of berberine, a bioactive compound richly encountered in the rhizome of Coptidis japonica, in TNBS-induced C3H/HeN and C3H/HeJ mice models of colitis and found that the treated mice presented deactivation of the NF-kB due to reductions in the TLR4 expression, NF-kB phosphorylation, and nuclear translocation, as well as increases in antioxidant SOD and Catalase (CAT) and anti-inflammatory IL-10 expressions, in addition to decreases in pro-inflammatory TNF-α, IL-1β, IL-6, iNOS, and COX-2 expressions.
- 4.5.11. Eriodictyol
- 4.5.12. Betulin
- 4.5.13. Naringin
- 4.5.14. 5-Hydroxy-4-methoxycanthin-6-one
- 4.5.15. Geniposide
- 4.5.16. Sesamin
- 4.5.17. Taxifolin
Hou et al. [49] showed that taxifolin, a natural antioxidant polyphenol with various bioactivities that is extracted from artichoke, onions, olive oil, grapes, milk thistle, citrus fruits and sorghum grain, could decrease NF-kB activation due to a reduced TNF-α, IL-1β and IL-6 expression levels, elevated Immunoglobulin A (SIgA), IL-10 and SOD expression levels and decreased p-NF-kB-p65 and p-IkBα in DSS-induced model of colitis.
- 4.5.18. Isobavachalcone
- 4.5.19. d-Pinitol
- 4.5.20. Paeoniflorin-6 -O-benzene Sulfonate
- 4.5.21. Thymol
- 4.5.22. Tricin
- 4.5.23. Aesculin
- 4.5.24. Ginsenoside Rk3
- 4.5.25. Lancemaside A
Joh et al. [57] evaluated the anti-inflammatory effects of lancemaside A, the main compound of the rhizome of Codonopsis lanceolata. In vitro, the treatment decreased this
activation of LR4-linked NF-kB. In vivo, there was an improvement in the thickening, shortening, ulceration, edema, and inflammation of the colon, regulated by the elevation of MPO, IL-6 mRNA, IL-1β mRNA, TNF-α mRNA, TLR4 mRNA, NF-kB (p-p65), and COX-2.
- 4.5.26. Tetramethylpyrazine
- 4.5.27. Daurisoline
- 4.5.28. Tetrandrine
- 4.5.29. Diosgenin
- 4.5.30. Mangiferin
Jeong et al. [62] performed in vivo and in vitro analyses to evaluate the antiinflammatory mechanisms of mangiferin, a compound found in plants such as Mangifera indica L., Anemarrhena asphodeloides and Cyclopia intermedia. In vitro, there was an increase in interleukin 1 receptor associated kinase 1 (IRAK1) phosphorylation and degradation, degradation of Interleukin 1 (IL-1) receptor-associated kinases (IRAK) 1, 2, and 4, increase in NF-kB activation, in transforming growth factor-β-activated kinase 1 (TAK1) phosphorylation and degradation, in IKKβ phosphorylation, in IκBα phosphorylation and degradation, in mediators inflammatory (PGE2, TNF-α expression, IL-1β expression, IL-6 expression, and COX-2), anti-inflammatory (IL-10 expression) and ROS (iNOS expression and NO). After treatment, there was a reversal of this scenario and a decrease in p65 translocation, MAPK p38 phosphorylation, ERK phosphorylation, and JNK phosphorylation. In vivo, before treatment, there was activation of IRAK1 and IKKβ and NF-kB; there was also an elevation of COX-2, iNOS, TNF-α, IL-1β, and IL-6. These conditions were suppressed by mangiferin.
- 4.5.31. Tryptanthrin
- 4.5.32. l-Theanine
- 4.5.33. Koreanaside A
- 4.5.34. 6-Gingerol
- 4.5.35. Lycopene
- 4.5.36. α-Mangostin
You et al. [68] used α-mangostin, the main xanthone derived from Garcinia mangostana L. plant. After induction of colitis by DSS, the animals presented weight loss, diarrhea,
intestinal bleeding, colon shrinkage, ulceration, erosion, crypt distortion, edema, and inflammation characterized by cellular infiltration. In addition, there was an increase in MPO, phosphorylation of IKKα, IκBα, ERK1/2, stress-activated protein kinases (SAPK/JNK), and p38, and activation of NF-kB and MAPK. After treatment with α-mangostin, the histopathological findings were reversed, and the regulation of NF-kB activation factors attenuated the pro-inflammatory pathways.
- 4.5.37. Ophiopogonin D
- 4.5.38. Alantolactone
- 4.5.39. Sinomenine
- 4.5.40. Convallatoxin
- 4.5.41. Fisetin
Sahu et al. [73] used fisetin, a flavonoid found in various fruits and vegetables (persimmons, mangoes, grapes, apples, strawberries, peaches, cucumbers, onions, and tomatoes),
in DSS-induced colitis in mice and LPS-induced cells. In vitro, there was an increase in nitrites, inflammatory cytokines (TNF-α, IL-1β, and IL-6), COX-2, iNOS, NF-kB-p65 nuclear translocation, IkBα phosphorylation, and degradation. After treatment, there was a reversal of this scenario. In vivo, before treatment, there was a decrease in weight and size of the colon, loss of crypts and goblet cells, and inflammation marked by infiltration, elevation of MPO, TNF-α, IL-1β, IL-6, nitrites, COX-2, iNOS, nuclear NF-kB (p65), phosphorylation of IκBα (p-IκBα/IκBα), NF-kB (p65)-DNA binding activity, p-p38/p38, p-ERK/ERK, Akt phosphorylation, and thiobarbituric acid reactive substances (TBARS), and GSH reduction. After treatment, there was a reversal of the histological and inflammatory pattern; however, p-ERK/ERK remained high.
- 4.5.42. Genipin
- 4.5.43. Piperine
- 4.5.44. Ligustilide
- 4.5.45. Evodiamine
- 4.5.46. Chrysin
Dou et al. [78] studied the effects of chrysin, a flavonoid found in many plant extracts, honey, and propolis, in a TNBS-induced C57BL/6 mice model of colitis. The control mice presented weight loss, diarrhea, fecal bleeding, crypt distortion, and inflammatory
exudate in the intestine. Additionally, the controls expressed elevated p-65, IkBα phosphorylation and degradation, NF-kB nuclear translocation, iNOS mRNA, ICAM-1 mRNA, MCP-1 mRNA, COX-2 mRNA, TNF-α mRNA, IL-6 mRNA, and MPO. The treated mice showed reversed inflammatory pattern through decreased p-65, IkBα phosphorylation and degradation, and NF-kB nuclear translocation, as well as by diminished expression of iNOS mRNA, ICAM-1 mRNA, MCP-1 mRNA, COX-2 mRNA, TNF-α mRNA, IL-6 mRNA, and MPO.
- 4.5.47. Wogonoside
- 4.5.48. Oxymatrine
- 4.5.49. Epicatechin
- 4.5.50. Thymoquinone
- 4.5.51. Fraxinellone
Wu et al. [83] investigated the effects of fraxinellone, a naturally occurring lactone (Dictamnus dasycarpus), on the NF-kB pathway and NLRP3 inflammasome in a model of colitis. In vitro, after treatment, there was a reduction in IL-1β and IL-18 expression, phosphorylation of IKKα/β, IκBα, p65 phosphorylation, p65, Caspase-1 activation, and NLRP3 inflammasome. After induction of colitis by DSS, there was weight loss, diarrhea, fecal bleeding, increased mortality, ulcers, loss of colon, and inflammation configured by cellular infiltration, the elevation of pro-inflammatory cytokines (IL-1β, IL-18, TNF-α, and IL-6) and NO. However, treatment with fraxinellone reversed the morphological lesions and reduced MPO, inflammatory cytokines, and expression of VCAM1, iNOS, and COX-2, in addition to increasing glutathione.
- 4.5.52. Artesunate
- 4.5.53. Aesculetin
- 4.5.54. Euphol
- 4.5.55. Nobiletin
- 4.5.56. Galangin
Gerges et al. [88] used galangin, a flavonol present in different types of propolis, such as Egyptian propolis and Alpinia officinarum, to evaluate its therapeutic potential in an in vivo model of colitis. The results showed that after the induction of colitis by DSS in mice, the animals showed an increase in ulceration, necrosis, and intestinal inflammation marked by cell infiltration, increased expression of TLR4, and High mobility group box 1 (HMGB1),
MDA, nuclear NF-kB-p65 and pro-inflammatory cytokines (TNF-α and IL-6), in addition to the reduction of antioxidant species (GSH). However, treatment with galangin reversed this inflammatory condition by inhibiting NF-kB activation, interfering with NF-kB-p65 translocation.
5. Conclusions
In summary, IBD are very common diseases associated primarily with a massive activation of the NF-kB pathways even during their first stages, positively affecting the disease progression. Nowadays, the clinical goal of regulation of NF-kB during IBD by inhibiting its activation is a field of study. However, many side effects derived from synthetic anti-inflammatory drugs and pharmacies can damage other organs and systems than the gastrointestinal. As an important source of innovative treatments against IBDrelated NF-kB activation, phytochemicals with biological activities regulating NF-kB have been explored and investigated. They are derived from medicinal plants and do not lead to serious adverse effects. The modulatory mechanisms involved in NF-kB regulation by phytochemicals result in decreased levels of TNF-α, IL-1β, IL-6, IFN-γ, and COX-2, and augmented occludin, claudin-1, zonula occludens-1, and IL-10 expression. Thus, further research should focus on the efficacy and safety of phytochemicals and other natural regulators of NF-kB on IBD.
Author Contributions: Conceptualization, L.F.L., A.R.d.O.d.S., M.D.B., E.L.G., A.C.A. and S.M.B.; methodology, L.F.L., A.R.d.O.d.S., M.D.B., E.L.G., A.C.A. and S.M.B.; validation, L.F.L., A.R.d.O.d.S., M.D.B., E.L.G., A.C.A. and S.M.B.; formal analysis, L.F.L., A.R.d.O.d.S., M.D.B., E.L.G., A.C.A. and S.M.B.; investigation, L.F.L., A.R.d.O.d.S., M.D.B., A.C.A.d.C., A.C.A. and S.M.B.; writingoriginal draft preparation, L.F.L., A.R.d.O.d.S., M.D.B., E.L.G., R.d.A.G., R.V.S., A.C.A. and S.M.B.; writing—review and editing, L.F.L., A.R.d.O.d.S., M.D.B., E.L.G., R.d.A.G., R.V.S., A.C.A. and S.M.B.; visualization, L.F.L., A.R.d.O.d.S., M.D.B., E.L.G., R.d.A.G., R.V.S., A.C.A. and S.M.B.; supervision, L.F.L., A.R.d.O.d.S., M.D.B., E.L.G., R.d.A.G., R.V.S., A.C.A. and S.M.B.; project administration, L.F.L., A.R.d.O.d.S., M.D.B., E.L.G., R.d.A.G., R.V.S., A.C.A. and S.M.B. All authors have read and agreed to the published version of the manuscript.
Funding: This research received no external funding.
Acknowledgments: The authors attribute Smart Servier (https://smart.servier.com/, accessed on 1 December 2022) for scientific images that were used in this article under an attribution license of public copyrights (https://creativecommons.org/licenses/by/3.0/, accessed on 1 December 2022) and under the disclaimer of warranties. All Smart Servier’s images were not changed in the writing of this article.
Conflicts of Interest: The authors declare no conflict of interest.
Figures
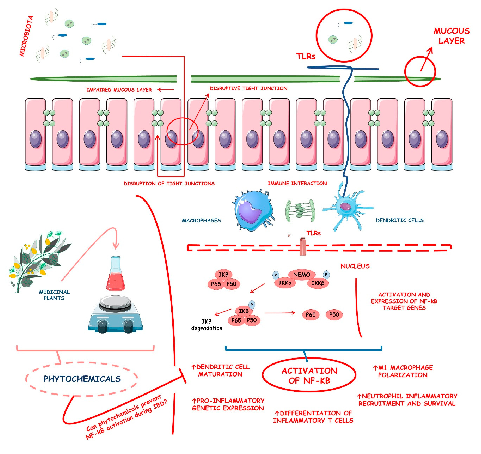
Figure 6
Inflammatory bowel diseases are associated with activation of the NF-kB signaling pathway. This diagram illustrates how various phytochemicals may counteract NF-kB pathway activation through inhibition of Toll-like receptors, IKK complex, and downstream transcriptional targets in IBD.
diagram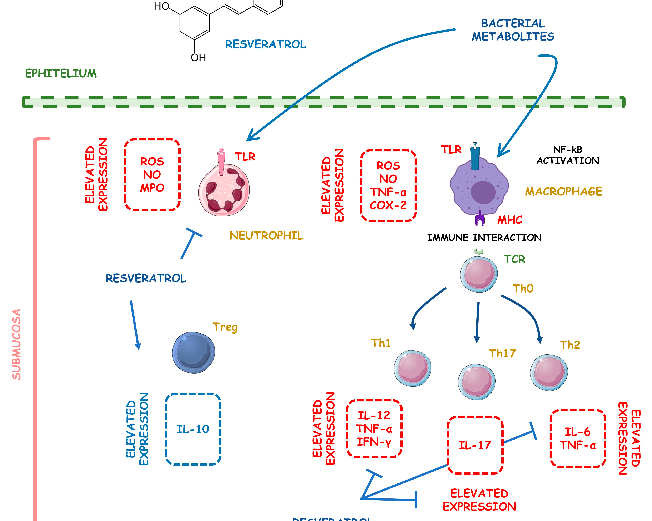
Figure 218
Supplementary figure from a comprehensive review of phytochemical effects on NF-kB regulation in inflammatory bowel disease. The review examines multiple plant-derived compounds with potential to attenuate chronic intestinal inflammation through modulation of key signaling pathways.
diagramUsed In Evidence Reviews
Similar Papers
The AAPS journal · 2013
Therapeutic roles of curcumin: lessons learned from clinical trials.
Digestive diseases and sciences · 2005
Curcumin therapy in inflammatory bowel disease: a pilot study.
Molecular nutrition & food research · 2013
Multitargeting by turmeric, the golden spice: From kitchen to clinic.
Gastroenterology · 2017
Diet as a Trigger or Therapy for Inflammatory Bowel Diseases.
Frontiers in endocrinology · 2023
Oxidative stress, hormones, and effects of natural antioxidants on intestinal inflammation in inflammatory bowel disease.
Microorganisms · 2022